Detecting clinically actionable somatic structural aberrations from targeted sequencing data
•Als PPTX, PDF herunterladen•
3 gefällt mir•562 views
Structural aberrations including deletions, insertions, inversions, tandem duplications, translocations, and more complex rearrangements constitute a frequent type of alteration in human tumors. Here, we sought to explore the potential to discover such events from targeted DNA sequence data in our CLIA-compliant molecular diagnostics laboratory. To detect somatic structural aberrations in individual tumors, we have developed an analytic framework in Perl & Python to detect these events in data generated by a hybridization capture-based, targeted sequencing clinical assay (MSK-IMPACT), which can reveal structural rearrangements as small as 500bp.
Melden
Teilen
Melden
Teilen
Empfohlen
Empfohlen
Weitere ähnliche Inhalte
Was ist angesagt?
Was ist angesagt? (19)
Reconstruction and analysis of cancerspecific Gene regulatory networks from G...

Reconstruction and analysis of cancerspecific Gene regulatory networks from G...
Purdue cancer center retreat poster Christy Cooper 12062014FINAL

Purdue cancer center retreat poster Christy Cooper 12062014FINAL
Gene expression profile analysis of human hepatocellular carcinoma using sage...

Gene expression profile analysis of human hepatocellular carcinoma using sage...
APPLICATION OF NEXT GENERATION SEQUENCING (NGS) IN CANCER TREATMENT

APPLICATION OF NEXT GENERATION SEQUENCING (NGS) IN CANCER TREATMENT
Proteogenomic analysis of human colon cancer reveals new therapeutic opportun...

Proteogenomic analysis of human colon cancer reveals new therapeutic opportun...
Abdulla, ICBO2011, Composite annotation for heart development

Abdulla, ICBO2011, Composite annotation for heart development
Bioinformatic Analysis of Synthetic Lethality in Breast Cancer

Bioinformatic Analysis of Synthetic Lethality in Breast Cancer
Genes and Tissue Culture Technology Assignment (G6)

Genes and Tissue Culture Technology Assignment (G6)
Andere mochten auch
Andere mochten auch (11)
Target Enrichment with NGS: Cardiomyopathy as a case study - BMR Genomics

Target Enrichment with NGS: Cardiomyopathy as a case study - BMR Genomics
Bioinformatica: Esercizi su Perl, espressioni regolari e altre amenità (BMR G...

Bioinformatica: Esercizi su Perl, espressioni regolari e altre amenità (BMR G...
Custom Enrichment Panels for Targeted Next Generation Sequencing

Custom Enrichment Panels for Targeted Next Generation Sequencing
GIAB Sep2016 Lightning megan cleveland targeted seq

GIAB Sep2016 Lightning megan cleveland targeted seq
Discovery and annotation of variants by exome analysis using NGS

Discovery and annotation of variants by exome analysis using NGS
Exome sequencing for disease gene identification and patient diagnostics, Gen...

Exome sequencing for disease gene identification and patient diagnostics, Gen...
Ähnlich wie Detecting clinically actionable somatic structural aberrations from targeted sequencing data
Genomics provides opportunities to develop precise tests for diagnostics, therapy selection and monitoring. From analyses of our studies and those of published results, 32 candidate genes were identified, whose expression appears related to clinical outcome of breast cancer. Expression of these genes was validated by qPCR and correlated with clinical follow-up to identify a gene subset for development of a prognostic test.Interrogating differences in expression of targeted gene sets to predict brea...

Interrogating differences in expression of targeted gene sets to predict brea...Enrique Moreno Gonzalez
Glioblastoma (GBM) is the most malignant brain tumor. Many abnormal secretion and
expression of cytokines have been found in GBM, initially speculated that the occurrence of
GBM may be involved in these abnormal secretion of cytokines. This study aims to detect the
association of cytokine genes with GBM.Genetic association between selected cytokine genes and glioblastoma in the H...

Genetic association between selected cytokine genes and glioblastoma in the H...Enrique Moreno Gonzalez
Ähnlich wie Detecting clinically actionable somatic structural aberrations from targeted sequencing data (20)
Bioinformatics-driven discovery of EGFR mutant Lung Cancer

Bioinformatics-driven discovery of EGFR mutant Lung Cancer
DNA Amplification is a Ubiquitous Mechanism of Oncogene Activation in Lung an...

DNA Amplification is a Ubiquitous Mechanism of Oncogene Activation in Lung an...
Gene expression profiling reveals molecularly and clinically distinct subtype...

Gene expression profiling reveals molecularly and clinically distinct subtype...
Sex-Based Difference in Gene Alterations and Biomarkers in Anal Squamous Cell...

Sex-Based Difference in Gene Alterations and Biomarkers in Anal Squamous Cell...
Oncomine Cancer Research Panel (OCP) | ESHG 2015 Poster PS12.131

Oncomine Cancer Research Panel (OCP) | ESHG 2015 Poster PS12.131
Next Generation Sequencing and its Applications in Medical Research - Frances...

Next Generation Sequencing and its Applications in Medical Research - Frances...
Assessing the clinical utility of cancer genomic and proteomic data across tu...

Assessing the clinical utility of cancer genomic and proteomic data across tu...
Interrogating differences in expression of targeted gene sets to predict brea...

Interrogating differences in expression of targeted gene sets to predict brea...
Genetic association between selected cytokine genes and glioblastoma in the H...

Genetic association between selected cytokine genes and glioblastoma in the H...
Kürzlich hochgeladen
Kürzlich hochgeladen (20)
Call Girls Shahdol Just Call 8250077686 Top Class Call Girl Service Available

Call Girls Shahdol Just Call 8250077686 Top Class Call Girl Service Available
7 steps How to prevent Thalassemia : Dr Sharda Jain & Vandana Gupta

7 steps How to prevent Thalassemia : Dr Sharda Jain & Vandana Gupta
Kolkata Call Girls Shobhabazar 💯Call Us 🔝 8005736733 🔝 💃 Top Class Call Gir...

Kolkata Call Girls Shobhabazar 💯Call Us 🔝 8005736733 🔝 💃 Top Class Call Gir...
Pune Call Girl Service 📞9xx000xx09📞Just Call Divya📲 Call Girl In Pune No💰Adva...

Pune Call Girl Service 📞9xx000xx09📞Just Call Divya📲 Call Girl In Pune No💰Adva...
Gorgeous Call Girls Dehradun {8854095900} ❤️VVIP ROCKY Call Girls in Dehradun...

Gorgeous Call Girls Dehradun {8854095900} ❤️VVIP ROCKY Call Girls in Dehradun...
Jual Obat Aborsi Di Dubai UAE Wa 0838-4800-7379 Obat Penggugur Kandungan Cytotec

Jual Obat Aborsi Di Dubai UAE Wa 0838-4800-7379 Obat Penggugur Kandungan Cytotec
👉Chandigarh Call Girl Service📲Niamh 8868886958 📲Book 24hours Now📲👉Sexy Call G...

👉Chandigarh Call Girl Service📲Niamh 8868886958 📲Book 24hours Now📲👉Sexy Call G...
Chandigarh Call Girls Service ❤️🍑 9809698092 👄🫦Independent Escort Service Cha...

Chandigarh Call Girls Service ❤️🍑 9809698092 👄🫦Independent Escort Service Cha...
💰Call Girl In Bangalore☎️63788-78445💰 Call Girl service in Bangalore☎️Bangalo...

💰Call Girl In Bangalore☎️63788-78445💰 Call Girl service in Bangalore☎️Bangalo...
💚Chandigarh Call Girls Service 💯Piya 📲🔝8868886958🔝Call Girls In Chandigarh No...

💚Chandigarh Call Girls Service 💯Piya 📲🔝8868886958🔝Call Girls In Chandigarh No...
Bandra East [ best call girls in Mumbai Get 50% Off On VIP Escorts Service 90...

Bandra East [ best call girls in Mumbai Get 50% Off On VIP Escorts Service 90...
🚺LEELA JOSHI WhatsApp Number +91-9930245274 ✔ Unsatisfied Bhabhi Call Girls T...

🚺LEELA JOSHI WhatsApp Number +91-9930245274 ✔ Unsatisfied Bhabhi Call Girls T...
Premium Call Girls Dehradun {8854095900} ❤️VVIP ANJU Call Girls in Dehradun U...

Premium Call Girls Dehradun {8854095900} ❤️VVIP ANJU Call Girls in Dehradun U...
Premium Call Girls Nagpur {9xx000xx09} ❤️VVIP POOJA Call Girls in Nagpur Maha...

Premium Call Girls Nagpur {9xx000xx09} ❤️VVIP POOJA Call Girls in Nagpur Maha...
👉 Chennai Sexy Aunty’s WhatsApp Number 👉📞 7427069034 👉📞 Just📲 Call Ruhi Colle...

👉 Chennai Sexy Aunty’s WhatsApp Number 👉📞 7427069034 👉📞 Just📲 Call Ruhi Colle...
Circulatory Shock, types and stages, compensatory mechanisms

Circulatory Shock, types and stages, compensatory mechanisms
Dehradun Call Girl Service ❤️🍑 8854095900 👄🫦Independent Escort Service Dehradun

Dehradun Call Girl Service ❤️🍑 8854095900 👄🫦Independent Escort Service Dehradun
Detecting clinically actionable somatic structural aberrations from targeted sequencing data
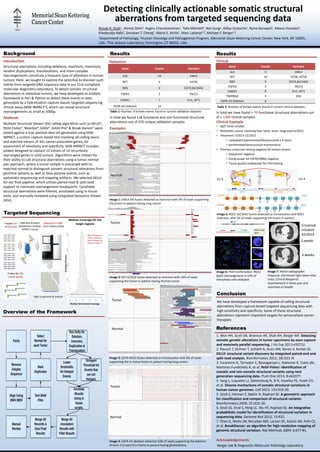
- 1. Detecting clinically actionable somatic structural aberrations from targeted sequencing data Ronak H. Shah1, Ahmet Zehir1, Raghu Chandramohan1, Talia Mitchell3, Wei Song1, Alifya Oultache1, Ryma Benayed1, Meera Hameed1, Khedoudja Nafa1, Donavan T. Cheng1, Maria E. Arcila1, Marc Ladanyi1,2, Michael F. Berger1,2 1Department of Pathology, 2Human Oncology and Pathogenesis Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA, 3The Jackson Laboratory, Farmington, CT 06032, USA Background Results Structural aberrations including deletions, insertions, inversions, tandem duplications, translocations, and more complex rearrangements constitute a frequent type of alteration in human tumors. Here, we sought to explore the potential to discover such events from targeted DNA sequence data in our CLIA-compliant molecular diagnostics laboratory. To detect somatic structural aberrations in individual tumors, we have developed an analytic framework in Perl & Python to detect these events in data generated by a hybridization capture-based, targeted sequencing clinical assay (MSK-IMPACT1), which can reveal structural rearrangements as small as 500bp. Multiple Structural Variant (SV) calling algorithms such as DELLY2, PeSV-Fisher3, Meerkat4, GASV5, GASV-Pro6 & Break-Dancer7 were tested against a true positive data set generated using MSK-IMPACT, a custom capture-based test involving all coding exons and selected introns of 341 cancer associated genes, for assessment of sensitivity and specificity. MSK-IMPACT includes probes designed to capture 33 introns of 14 recurrently rearranged genes in solid tumors. Algorithms were chosen for their ability to call structural aberrations using a tumor-normal pair approach, where a tumor sample is processed with its matched normal to distinguish somatic structural alterations from germline variants as well as false positive events, such as systematic sequencing and mapping artifacts. We selected DELLY for our final pipeline, which utilizes paired-read & split-read support to nominate rearrangement breakpoints. Candidate structural aberrations were filtered, annotated using in-house tools, and manually reviewed using Integrated Genomics Viewer (IGV). Targeted Sequencing Results translocatio n translocatio n chr 6 chr 4 SLC34A2 ROS1 SLC34A2 30 31 32 ROS1 Image 5: ROS1-SLC34A2 fusion detected as translocation and ROS1 inversion, with 3% of reads supporting the fusion in patient. Conclusion Hybridize & select (NimbleGen SeqCap: IMPACT Assay) Overview of the Framework Crizotinib initiated 02/2014 We have developed a framework capable of calling structural aberrations from capture-based targeted sequencing data with high sensitivity and specificity. Some of these structural aberrations represent important targets for personalized cancer therapies. 1. Won HH, Scott SN, Brannon AR, Shah RH, Berger MF. Detecting somatic genetic alterations in tumor specimens by exon capture and massively parallel sequencing. J Vis Exp 2013:e50710. 2. Rausch T, Zichner T, Schlattl A, Stutz AM, Benes V, Korbel JO. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 2012; 28:i333-i9. 3. Escaramis G, Tornador C, Bassaganyas L, Rabionet R, Tubio JM, Martinez-Fundichely A, et al. PeSV-Fisher: identification of somatic and non-somatic structural variants using next generation sequencing data. PLoS One 2013; 8:e63377. 4. Yang L, Luquette LJ, Gehlenborg N, Xi R, Haseley PS, Hsieh CH, et al. Diverse mechanisms of somatic structural variations in human cancer genomes. Cell 2013; 153:919-29. 5. Sindi S, Helman E, Bashir A, Raphael BJ. A geometric approach for classification and comparison of structural variants. Bioinformatics 2009; 25:i222-30. 6. Sindi SS, Onal S, Peng LC, Wu HT, Raphael BJ. An integrative probabilistic model for identification of structural variation in sequencing data. Genome Biol 2012; 13:R22. 7. Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods 2009; 6:677-81. Acknowledgements Introduction Methods Prepare 24- 48 libraries Probes for 341 cancer genes Sequence to 500- 1000X (HiSeq 2500) Align to genome & analyze 98% of targets at >50% of median 99% of targets at >20% of median References Berger Lab & Diagnostic Molecular Pathology Laboratory Validation Gene Events Partners ALK 14 EML4 RET 4 KIF5B ROS 3 CD74,SLC34A2 FGFR3 2 TACC3 EWSR1 7 FLI1, WT1 EGFR vIII Deletion 14 In total we found 118 functional and non-functional structural aberrations out of 270 unique validation samples. Examples Tumor Normal Image 1: EML4-Alk fusion detected as inversion with 3% of reads supporting the fusion in patient having lung cancer. Tumor Normal Image 2: RET-CCDC6 fusion detected as inversion with 10% of reads supporting the fusion in patient having thyroid cancer. Tumor Normal Image 3: CD74-ROS1 fusion detected as translocation with 5% of reads supporting the in-frame fusion in patient having lung cancer. Tumor Normal Image 4: EGFR vIII deletion detected 10% of reads supporting the deletion of exon 2 to exon 8 in-frame in patient having glioblastoma. Clinical Gene Events Partners ALK 3 EML4 RET 10 CCD6, KIF5B ROS 9 CD74,SLC34A2 FGFR3 2 TACC3 EWSR1 9 FLI1, WT1 TMPRSS2 5 ERG EGFR vIII Deletion 6 In total we have found > 70 functional structural aberrations out of > 1300 clinical samples. Clinical Example • 58/F never smoker • Metastatic cancer involving liver, bone, brain: diagnosed 6/2013 • Treatment 7/2013-12/2013 • carboplatin/pemetrexed/bevacizumab x 4 cycles • pemetrexed/bevacizumab maintenance • Previous molecular testing negative for known drivers • Sequenom negative • Sizing assays for EGFR/ERBB2 negative • Tissue quality inadequate for FISH testing inversion ROS1 SLC34A2 1 2 3 4 28 29 30 1 2 3 4 25 26 27 28 29 30 3’ probe ROS1 5’ probe ROS1 break apart Image 6: FISH Confirmation: ROS1 6q22 rearrangement in 54% of interphase cells analyzed 0 weeks 4 weeks Image 7: Minor radiographic response: decreased right lower lobe mass. Clinical Response: Improvement in bone pain and shortness of breath Table 1: Number of known events found in current validation datasets. Table 2: Number of known events found in current clinical datasets. Median Coverage for the target regions Median Normalized Coverage Fraction of Exons
