Aza Cope Rearrangement of Propargyl Enammonium Cations Catalyzed by a Self-Assembled "Nanozyme"
•Als PPT, PDF herunterladen•
1 gefällt mir•1,235 views
Melden
Teilen
Melden
Teilen
Empfohlen
Empfohlen
Weitere ähnliche Inhalte
Was ist angesagt?
Was ist angesagt? (20)
B.tech. ii engineering chemistry unit 4 B organic chemistry

B.tech. ii engineering chemistry unit 4 B organic chemistry
Ähnlich wie Aza Cope Rearrangement of Propargyl Enammonium Cations Catalyzed by a Self-Assembled "Nanozyme"
Understanding ionic structure and dynamics in novel electrolytes; Paving the way to improved energy storage by Maria ForsythUnderstanding ionic structure and dynamics in novel electrolytes; Paving the ...

Understanding ionic structure and dynamics in novel electrolytes; Paving the ...Trinity College Dublin
Ähnlich wie Aza Cope Rearrangement of Propargyl Enammonium Cations Catalyzed by a Self-Assembled "Nanozyme" (20)
Cyclic Voltammetry: Principle, Instrumentation & Applications

Cyclic Voltammetry: Principle, Instrumentation & Applications
Understanding ionic structure and dynamics in novel electrolytes; Paving the ...

Understanding ionic structure and dynamics in novel electrolytes; Paving the ...
Simulating Membrane Channels, E. Tajkhorshid, Part 2

Simulating Membrane Channels, E. Tajkhorshid, Part 2
Interpretation Of Organic Molecules By Mass Spectra

Interpretation Of Organic Molecules By Mass Spectra
Nucleation and Growth of Crystal Phases from Amorphous Solids

Nucleation and Growth of Crystal Phases from Amorphous Solids
Heterogeneous Catalyst-opportunity and challenges.ppt

Heterogeneous Catalyst-opportunity and challenges.ppt
Heterogeneous Catalysis - Opportunities and Challenges

Heterogeneous Catalysis - Opportunities and Challenges
9. m. pharm interpretation of 1 h nmr jntu pharmacy

9. m. pharm interpretation of 1 h nmr jntu pharmacy
Mehr von Matthew MacLennan
Mehr von Matthew MacLennan (12)
A brand new method of analyzing organic contaminants in oil sands tailings po...

A brand new method of analyzing organic contaminants in oil sands tailings po...
The Emergence of New Radical-Cationic Amino Acid Dynamics: The Proton Patches...

The Emergence of New Radical-Cationic Amino Acid Dynamics: The Proton Patches...
20140127 matthew - literature presentation - lipids

20140127 matthew - literature presentation - lipids
2013 02 12_matthew s. mac_lennan_ink ageing chemistry

2013 02 12_matthew s. mac_lennan_ink ageing chemistry
2013 10 21_matthew s. mac_lennan_marshmallow polymer synthesis

2013 10 21_matthew s. mac_lennan_marshmallow polymer synthesis
Chansons d'acadie 1ère série père anselme-frère daniel

Chansons d'acadie 1ère série père anselme-frère daniel
Kürzlich hochgeladen
Kürzlich hochgeladen (20)
Creating and Analyzing Definitive Screening Designs

Creating and Analyzing Definitive Screening Designs
Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...

Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...
Forensic Biology & Its biological significance.pdf

Forensic Biology & Its biological significance.pdf
PossibleEoarcheanRecordsoftheGeomagneticFieldPreservedintheIsuaSupracrustalBe...

PossibleEoarcheanRecordsoftheGeomagneticFieldPreservedintheIsuaSupracrustalBe...
Recombinant DNA technology (Immunological screening)

Recombinant DNA technology (Immunological screening)
Biopesticide (2).pptx .This slides helps to know the different types of biop...

Biopesticide (2).pptx .This slides helps to know the different types of biop...
All-domain Anomaly Resolution Office U.S. Department of Defense (U) Case: “Eg...

All-domain Anomaly Resolution Office U.S. Department of Defense (U) Case: “Eg...
SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE

SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE
Pests of mustard_Identification_Management_Dr.UPR.pdf

Pests of mustard_Identification_Management_Dr.UPR.pdf
Stunning ➥8448380779▻ Call Girls In Panchshil Enclave Delhi NCR

Stunning ➥8448380779▻ Call Girls In Panchshil Enclave Delhi NCR
Kochi ❤CALL GIRL 84099*07087 ❤CALL GIRLS IN Kochi ESCORT SERVICE❤CALL GIRL

Kochi ❤CALL GIRL 84099*07087 ❤CALL GIRLS IN Kochi ESCORT SERVICE❤CALL GIRL
TEST BANK For Radiologic Science for Technologists, 12th Edition by Stewart C...

TEST BANK For Radiologic Science for Technologists, 12th Edition by Stewart C...
Hubble Asteroid Hunter III. Physical properties of newly found asteroids

Hubble Asteroid Hunter III. Physical properties of newly found asteroids
Aza Cope Rearrangement of Propargyl Enammonium Cations Catalyzed by a Self-Assembled "Nanozyme"
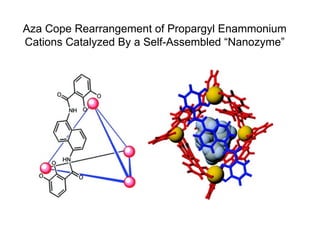
- 1. Aza Cope Rearrangement of Propargyl Enammonium Cations Catalyzed By a Self-Assembled “Nanozyme”
- 2. Dr. Kenneth Raymond Born 1942; B. A. Reed College (1964); Ph. D. Northwestern University (1968); Alfred P. Sloan Research Fellow (1971- 1973); Miller Research Professor (1977-1978, 1996, 2004); Guggenheim Fellow (1980-1981); Selected as one of the "Technology 100, 1981" by Technology Magazine; American Association for the Advancement of Science Fellow (1984); DOE Ernest O. Lawrence Award (1984); Lawrence Berkeley Laboratory Technology Transfer award (1988, 1991); Humboldt Research Award for Senior U.S. Scientists (1992); American Chemical Society Alfred Bader Award in Bioinorganic or Bioorganic Chemistry (1994); Erskine Fellow, University of Canterbury, New Zealand (1997); Elected to National Academy of Sciences (1997); Basolo Medal, Northwestern University (1997); Max-Planck-Institut fur Strahlenchemie "Frontiers in Biological Chemistry" Award (1997); Elected to the American Academy of Arts and Sciences (2001); Reed College Howard Vollum Award (2002); ACS Auburn Section G. M. Kosolapoff Award (2004); Izatt-Christensen Award in Macrocyclic Chemistry (2005); Joe L. Franklin Memorial Lectureship (2006); Paulo Fasella Lectureship (2006); UC Berkeley Chancellor's Professor, (2007-). 426 Pubs 126 Inorg Chem 113 JACS 21 Angew
- 3. The Cope Rearrangement • 3,3 sigmatropic rearrangement • Aza Cope rearrangement
- 4. The Catalyst • M4L6 12- assembly • M = Ga3+ (Al3+ , Fe3+ ); L = N,N’-bis(2,3-dihydroxybenzoyl)- 1,5-diaminonaphthalene • Racemic mixture of homochiral ΔΔΔΔ and ΛΛΛΛ enantiomers • Anionic nature makes soluble in H2O, with hydrophobic core • Water-labile cations (ketone-derived iminium ions, diazonium, tropylium, phosphine-acetone adducts) are encapsulated
- 6. Basic Cope rearrangement of enammonium
- 7. 1 H NMR • NMR in D2O • Hi-res ES-TOF MS in H2O
- 9. Rate constants Compound R kfree=(10-8 s-1 ) kencaps=(10-8 s-1 ) kencaps/kfree 2 H 62.4 237 4 3 Me 62.3 6200 100 4 Et 20.0 3670 184 5 n-Pr 19.5 1920 98 6 i-Pr 6.7 870 129 7 n-Bu 15.1 73 5 8 i-Bu 17.0 477 28 9 s-Bu 50.0 1150 23 R=Me has fastest encapsulation rate; zeroth order when >3 eq substrate; RLS = rearrangementrate depends on [host-bound substrate].
- 10. • Catalyzed reaction on the left; uncatalyzed on the right. ΔH‡ is more negative for the catalyzed reaction • Entropy for catalyzed reaction is >20 J/mol more positive than for uncatalyzed • Entropy-based rate increase R S h k RT H T k Brate ‡‡ lnln ∆ + + ∆− =
- 11. Michaelis-Menten in the house Vmax = 1.2 x 10-4 mM.s-1 Km = 0.67 mM kcat = 7.0 x 10-5 s-1 Vmax = ~1.05 x 10-4 mM.s-1 Km = >1.7 mM kcat = ? x 10-5 s-1
- 12. The Emergence of a New Radical-Cationic Amino Acid Dynamics: The Proton Patches Model Matthew MacLennan 1 J. Mol. Struct. THEOCHEM 0 Angew. Chem. 0 JACS
- 13. Dr. Galina Orlova Rostov University, Russia, 1981-1998 (R. Minyaev) Southern Illinois University at Carbondale 1996/97 (S. Scheiner) University at Guelph 1998/2002 (J. D. Goddard) York University 2002/2004 (K.W.M. Siu, D.K. Bohme, A.C. Hopkinson) 42 Pubs 10 J Phys Chem A 5 JACS
- 14. Methodology • Lowest energy conformers of neutral amino acid • Geometry in Gaussian • Charge = +1; Multiplicity = 2 (ionization) • Run CPMD simulation to test
- 15. “Proton Scissors” 12 AWFULLY COMPLICATED STEPS This fragmentation of C-N bond to give oxazolone cation and neutral fragment occurs at 31.3 kcal/mol. This size barrier is common with protonated species (between 30 and 40 kcal/mol). The fragmentation of any C-N or C-C bond in GGG is always preceded by proton transfer. Fragments + +
- 16. Amino Acids • Arginine • Asparigine • Aspartic Acid • Threonine • Tryptophan
- 18. Arg+• 0 2 4 6 8 10 12 0 100 200 300 400 500 600 700 800 Time (fs) C-CBondLength(Angstroms)
- 19. Arg+• 0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6 1.8 2 0 100 200 300 400 500 600 700 800 Time (fs) N-HBondDistance(Angstroms)
- 21. Asn+• 0 0.2 0.4 0.6 0.8 1 1.2 1.4 1.6 1.8 2 2.2 0 100 200 300 400 500 600 700 800 Time (fs) O-HBondDistance(Angstroms)
- 22. Asn+• 0 1 2 3 4 5 6 7 0 100 200 300 400 500 600 700 800 Time (fs) C-CBondDistance(Angstroms)
- 24. Asp+• 0 1 2 3 4 5 6 7 0 100 200 300 400 500 Time (fs) C-CBondDistance(Angstroms)
- 25. Asp+• 0 1 2 3 4 5 6 0 100 200 300 400 500 Time (fs) O-HBondDistance(Angstroms) ~124 fs
- 28. Summary • Radical-cationic amino acids do not obey the “proton scissors” motif (Proton transfer before C- C bond cleavage); we see variety • Arg+• , Asp+• , and Thr+• (conformer 2) show C-C bond cleavage before proton transfer • Asn+• shows C-C bond cleavage and proton transfer occurring almost simultaneously • Thr+• , Trp+• show C-C bond cleavage without any proton transfer • Explanation for lack of IE potentials of amino acids
