New Microsoft PowerPoint Presentation.pptx
•Als PPTX, PDF herunterladen•
0 gefällt mir•5 views
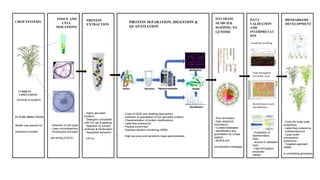
This document discusses current limitations and future directions in plant proteomics. It outlines several key challenges including the diversity of crop systems, limitations in tissue and cell isolations, difficulties in extracting and separating low abundant proteins, poor database annotation, and the need for improved validation and bioinformatics tools. Future work aims to address these issues through approaches like label-free proteomics, targeted assays, orthoproteomics, and large-scale comparative studies across genotypes.
Melden
Teilen
Melden
Teilen
Empfohlen
Empfohlen
Weitere ähnliche Inhalte
Ähnlich wie New Microsoft PowerPoint Presentation.pptx
Ähnlich wie New Microsoft PowerPoint Presentation.pptx (20)
Developing standards for metabonomics as a clinical tool.

Developing standards for metabonomics as a clinical tool.
Predictive Models for Mechanism of Action Classification from Phenotypic Assa...

Predictive Models for Mechanism of Action Classification from Phenotypic Assa...
Synthetic Biology Approaches to identify Enzymes from Metagenome by Maliha Ra...

Synthetic Biology Approaches to identify Enzymes from Metagenome by Maliha Ra...
Large-scale analysis of the yeast proteome by multi dimensional protein ident...

Large-scale analysis of the yeast proteome by multi dimensional protein ident...
Mehr von euphemism22
Mehr von euphemism22 (20)
EDITED Rev3-Low Light-Bases-L Behera-23Nov-23(AAU).pptx

EDITED Rev3-Low Light-Bases-L Behera-23Nov-23(AAU).pptx
Kürzlich hochgeladen
VVVIP Call Girls In Greater Kailash ➡️ Delhi ➡️ 9999965857 🚀 No Advance 24HRS Live
Booking Contact Details :-
WhatsApp Chat :- [+91-9999965857 ]
The Best Call Girls Delhi At Your Service
Russian Call Girls Delhi Doing anything intimate with can be a wonderful way to unwind from life's stresses, while having some fun. These girls specialize in providing sexual pleasure that will satisfy your fetishes; from tease and seduce their clients to keeping it all confidential - these services are also available both install and outcall, making them great additions for parties or business events alike. Their expert sex skills include deep penetration, oral sex, cum eating and cum eating - always respecting your wishes as part of the experience
(07-May-2024(PSS)VVVIP Call Girls In Greater Kailash ➡️ Delhi ➡️ 9999965857 🚀 No Advance 24HRS...

VVVIP Call Girls In Greater Kailash ➡️ Delhi ➡️ 9999965857 🚀 No Advance 24HRS...Call Girls In Delhi Whatsup 9873940964 Enjoy Unlimited Pleasure
Kürzlich hochgeladen (20)
7.pdf This presentation captures many uses and the significance of the number...

7.pdf This presentation captures many uses and the significance of the number...
Mysore Call Girls 8617370543 WhatsApp Number 24x7 Best Services

Mysore Call Girls 8617370543 WhatsApp Number 24x7 Best Services
Call Girls Hebbal Just Call 👗 7737669865 👗 Top Class Call Girl Service Bangalore

Call Girls Hebbal Just Call 👗 7737669865 👗 Top Class Call Girl Service Bangalore
Call Girls Electronic City Just Call 👗 7737669865 👗 Top Class Call Girl Servi...

Call Girls Electronic City Just Call 👗 7737669865 👗 Top Class Call Girl Servi...
Lucknow 💋 Escorts in Lucknow - 450+ Call Girl Cash Payment 8923113531 Neha Th...

Lucknow 💋 Escorts in Lucknow - 450+ Call Girl Cash Payment 8923113531 Neha Th...
Value Proposition canvas- Customer needs and pains

Value Proposition canvas- Customer needs and pains
Yaroslav Rozhankivskyy: Три складові і три передумови максимальної продуктивн...

Yaroslav Rozhankivskyy: Три складові і три передумови максимальної продуктивн...
Insurers' journeys to build a mastery in the IoT usage

Insurers' journeys to build a mastery in the IoT usage
Grateful 7 speech thanking everyone that has helped.pdf

Grateful 7 speech thanking everyone that has helped.pdf
VVVIP Call Girls In Greater Kailash ➡️ Delhi ➡️ 9999965857 🚀 No Advance 24HRS...

VVVIP Call Girls In Greater Kailash ➡️ Delhi ➡️ 9999965857 🚀 No Advance 24HRS...
Call Girls Navi Mumbai Just Call 9907093804 Top Class Call Girl Service Avail...

Call Girls Navi Mumbai Just Call 9907093804 Top Class Call Girl Service Avail...
0183760ssssssssssssssssssssssssssss00101011 (27).pdf

0183760ssssssssssssssssssssssssssss00101011 (27).pdf
Call Girls In DLf Gurgaon ➥99902@11544 ( Best price)100% Genuine Escort In 24...

Call Girls In DLf Gurgaon ➥99902@11544 ( Best price)100% Genuine Escort In 24...
Call Girls Pune Just Call 9907093804 Top Class Call Girl Service Available

Call Girls Pune Just Call 9907093804 Top Class Call Girl Service Available
New Microsoft PowerPoint Presentation.pptx
- 1. CURREN T LIMITATI ONS CROP SYSTEMS TISSUE AND CELL ISOLATIONS PROTEIN EXTRACTION PROTEIN SEPARATION, DIGESTION & QUANTITATION DATABASE SEARCH & MAPPING TO GENOME DATA VALIDATION AND INTERPRETAT ION metabolite profiling High throughput enzymatic assay Bioinformatics tools and databases BIOMARKERS DEVELOPMENT CURRENT LIMITATIONS FUTURE DIRECTIONS - Diversity of systems Model crop species for proteomics studies - Selection of cell types - Laser microdissection - Fluorescent-activated cell sorting (FACS) - Highly abundant proteins - Detergent compatible with 2-D gel & labelling - Depletion by solvent, antibody & fractionation - Sequential extraction - CPLLs - Costs of DIGE and labelling approaches - Detection & quantitation of low abundant proteins - Characterization of protein modifications - Label-free proteomics - Peptide enrichment - Selected reaction monitoring (SRM) - High accuracy and sensitivity mass spectrometers - Poor annotation - High sequence redundancy - Curated databases - Identification and quantitation by unique peptide - Multihit and normalization strategies - Availability of bioinformatics tools - Access to validation tools - High-throughput enzymatic assays - Costs for large scale proteomics - Label-free proteomics - Orthoproteomics - Large scale comparative proteomics - Targeted approach (SRM) in contrasting genotypes