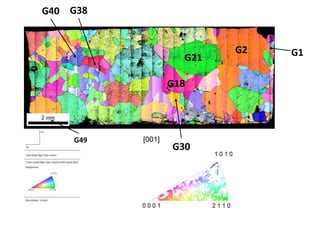
Ni pure mg corrected data
- 1. G2 G1 G21 G38 G49 G40 G18 G30
- 3. G4- Test #015 Estat = mean: 42.7524 median: 42.7659 stdev: 1.9003 min: 38.1723 max: 47.3500 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.1 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=38.9023; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 0 -1 -2 -3 -4 -5 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 4. G4- Test #015 100 50 0 30 35 40 45 50 Modulus [GPa] 200 100 0 0.97 0.98 0.99 1 1.01 R2 50 0 150 200 250 300 Modulus Length [#points] 100 50 0 -0.8 -0.7 -0.6 -0.5 -0.4 H r [nm] 100 50 2.4 2.6 2.8 3 -3 x 10 0 Fit1 Avg.Abs.Res. 7 8 9 10 11 -3 x 10 100 50 0 Fit1 Max.Abs.Res. 50 0 0.16 0.18 0.2 0.22 Fit2 Avg.Abs.Res. 0 0.5 1 1.5 100 50 0 Fit2 Max.Abs.Res. 100 50 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 100 50 0 P change [mN/mN] 50 0 0.009 0.01 0.011 0.012 Fit4 Avg.Abs.Res. 50 0 0.028 0.029 0.03 0.031 0.032 Fit4 Max.Abs.Res. 100 50 0 0.05 0.06 0.07 0.08 P* [mN] 100 50 0 11.5 12 12.5 13 13.5 h* [nm] 0 1 2 3 4 -4 x 10 40 20 0 dP [mN] 0 0.2 0.4 0.6 0.8 100 50 0 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.97;... 'R22', 0.97;... 'R23', 0.97;... 'seg_end', 700;... % 'AAR1', 0.003;... 'MAR1', 0.012;... % 'MAR2', 1.4;... 'MAR4', 0.032;... % 'AAR4', 0.02;... 'P*', 0.09;... 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dH', 0;... % 'h_change', [0.35 0.18];... 'dP', 0.0004};...
- 5. G7- Test #025 Estat = mean: 44.6889 median: 44.5730 stdev: 1.7866 min: 41.6071 max: 49.0945 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=40.9861; Bunge=0, 0, 0; Strength=0.77868; Hardening=1.0356 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data Yield Point 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 6 0 1 2 3 4 5 4 x 10 0 -0.5 -1 -1.5 -2 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 6. G7- Test #025 10 5 0 10 5 10 5 0 5 0 Filt ={ 'Modulus', [50 37];... 'R21', 0.984;... 'R22', 0.984;... 'R23', 0.984;... % 'AAR2', 0.4;... 'MAR1', 0.05;... 'MAR2', 5;... 'MAR4', 0.06;... % 'AAR4', 0.02;... 'P*', 0.4;... 'ModLength', 100;... % % % % 'h*', 95.5;... % % % 'AAR2', 0.22;... 'seg_end', 960;... 'dP', 0.0015};... 35 40 45 50 Modulus [GPa] 10 5 0 0.98 0.985 0.99 0.995 1 R2 10 5 0 120 140 160 180 Modulus Length [#points] 10 5 0 -1.2 -1.1 -1 -0.9 -0.8 H r [nm] 4 6 8 10 -3 x 10 0 Fit1 Avg.Abs.Res. 10 5 0 0.015 0.02 0.025 0.03 Fit1 Max.Abs.Res. 0.3 0.32 0.34 20 10 0 Fit2 Avg.Abs.Res. 1 1.2 1.4 1.6 1.8 10 5 0 Fit2 Max.Abs.Res. -0.1 0 0.1 0.2 0.3 h change [nm/nm] 0 0.05 0.1 10 5 0 P change [mN/mN] 10 5 0 0.011 0.012 0.013 0.014 0.015 Fit4 Avg.Abs.Res. 0.04 0.045 0.05 20 10 0 Fit4 Max.Abs.Res. 0.25 0.3 0.35 0.4 0.45 P* [mN] 5 0 29 30 31 32 33 h* [nm] 0 0.5 1 1.5 -3 x 10 10 5 0 dP [mN] 0 0.5 1 10 5 0 dH [nm] effective sample Fit1 Fit2 Fit3
- 7. G8- Test #032 Estat = mean: 36.1909 median: 36.1927 stdev: 0.5636 min: 35.0410 max: 38.2811 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=32.8002; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 200 400 600 10 8 6 4 2 0 -2 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 6 0 1 2 3 4 4 x 10 0 -0.5 -1 -1.5 -2 -2.5 -3 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 8. G8- Test #032 500 0 400 200 400 200 0 400 200 0 Filt ={ 'Modulus', [50 25];... 'R21', 0.97;... 'R22', 0.97;... 'R23', 0.97;... 'seg_end', 850;... 'AAR2', 0.22;... % 'MAR2', 0.65;... % % 'MAR2', 0.9;... 'MAR4', 0.027;... 'AAR4', 0.0098;... 'P*', 0.07;... 'ModLength', 150;... % % 'h*', 95.5;... % % 'AAR2', 0.22;... % % 'dH', 0;... % % 'h_change', [0.35 0.18];... 'dP', 0.0004};... 30 35 40 Modulus [GPa] 1000 500 0 0.985 0.99 0.995 1 1.005 R2 400 200 0 200 250 300 350 400 Modulus Length [#points] 400 200 0 -0.9 -0.8 -0.7 -0.6 H r [nm] 2.5 3 3.5 -3 x 10 0 Fit1 Avg.Abs.Res. 400 200 0 0.0115 0.012 0.0125 0.013 0.0135 Fit1 Max.Abs.Res. 400 200 0 0.185 0.19 0.195 0.2 0.205 Fit2 Avg.Abs.Res. 0.8 1 1.2 1.4 400 200 0 Fit2 Max.Abs.Res. -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 400 200 0 P change [mN/mN] 500 7.5 8 8.5 9 -3 x 10 0 Fit4 Avg.Abs.Res. 400 200 0 0.025 0.026 0.027 0.028 Fit4 Max.Abs.Res. 0.055 0.06 0.065 0.07 0.075 P* [mN] 500 0 94.4 94.6 94.8 95 95.2 h* [nm] 0 1 2 3 4 -4 x 10 200 100 0 dP [mN] 400 200 0 -0.4 -0.2 0 0.2 0.4 dH [nm] effective sample Fit1 Fit2 Fit3
- 9. G7- Test #066 Estat = mean: 39.0649 median: 38.7185 stdev: 2.3329 min: 34.6295 max: 46.6233 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=35.4797; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 0 -1 -2 -3 -4 -5 -6 -7 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 10. G7- Test #066 1000 500 0 2000 1000 1000 500 0 1000 500 0 Filt ={ 'Modulus', [50 25];... 'R21', 0.97;... 'R22', 0.97;... 'R23', 0.97;... 'seg_end', 800;... 'AAR2', 0.2;... % % 'MAR2', 0.65;... % % % 'MAR2', 0.9;... 'MAR4', 0.03;... % 'AAR4', 0.0098;... % 'P*', 0.07;... 'ModLength', 150;... % % 'h*', 95.5;... 'AAR2', 0.22;... % % 'dH', 0;... % % 'h_change', [0.35 0.18];... 'dP', 0.0004};... 30 35 40 45 50 Modulus [GPa] 2000 1000 0 0.97 0.98 0.99 1 1.01 R2 1000 500 0 100 200 300 400 Modulus Length [#points] 1000 500 0 -1.2 -1 -0.8 -0.6 -0.4 H r [nm] 2 3 4 5 -3 x 10 0 Fit1 Avg.Abs.Res. 2000 1000 0 0.005 0.01 0.015 0.02 0.025 Fit1 Max.Abs.Res. 0.16 0.18 0.2 1000 500 0 Fit2 Avg.Abs.Res. 1000 500 0 0.5 1 1.5 Fit2 Max.Abs.Res. -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 4 6 8 10 -3 x 10 1000 500 0 Fit4 Avg.Abs.Res. 1000 500 0 0.02 0.025 0.03 Fit4 Max.Abs.Res. 0.04 0.06 0.08 0.1 P* [mN] 1000 500 0 10 11 12 13 h* [nm] 0 1 2 3 4 -4 x 10 500 0 dP [mN] 1000 500 0 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3
- 11. G7- Test #084 Estat = mean: 37.3097 median: 37.3282 stdev: 0.4476 min: 36.2891 max: 38.3929 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=33.8199; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 1 0 -1 -2 -3 -4 -5 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 12. G7- Test #084 100 50 0 32 34 36 38 40 Modulus [GPa] 200 100 0 0.96 0.98 1 1.02 R2 100 50 0 300 350 400 450 Modulus Length [#points] 100 50 0 -0.9 -0.8 -0.7 -0.6 H r [nm] 200 100 3.2 3.4 3.6 3.8 4 -3 x 10 0 Fit1 Avg.Abs.Res. 500 0 0.012 0.013 0.014 0.015 0.016 Fit1 Max.Abs.Res. 100 50 0 0.18 0.19 0.2 0.21 Fit2 Avg.Abs.Res. 0.7 0.8 0.9 1 100 50 0 Fit2 Max.Abs.Res. 200 100 0 -0.1 0 0.1 0.2 0.3 h change [nm/nm] 0 0.05 0.1 0.15 0.2 200 100 0 P change [mN/mN] 100 50 0 0.01 0.0105 0.011 0.0115 0.012 Fit4 Avg.Abs.Res. 100 50 0 0.034 0.035 0.036 0.037 0.038 Fit4 Max.Abs.Res. 100 50 0 0.04 0.042 0.044 0.046 0.048 P* [mN] 7 7.5 8 8.5 100 50 0 h* [nm] 0 1 2 3 4 -4 x 10 100 50 0 dP [mN] 100 50 0 -0.4 -0.2 0 0.2 0.4 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.96;... 'R22', 0.96;... 'R23', 0.96;... 'seg_end', 770;... 'AAR2', 0.23;... % % % 'MAR2', 0.65;... 'MAR2', 1.1;... 'MAR4', 0.037;... % % 'AAR4', 0.0098;... 'P*', 0.048;... 'ModLength', 300;... % % % 'h*', 95.5;... % 'AAR2', 0.22;... % % % 'dH', 0;... % % % 'h_change', [0.35 0.18];... 'dP', 0.0004};...
- 13. G7- Test #091 Estat = mean: 39.9326 median: 40.1552 stdev: 1.2497 min: 36.3161 max: 42.2784 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.1 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=36.3565; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 1 0 -1 -2 -3 -4 -5 5 -1 0 1 2 3 4 4 x 10 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 0.1 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 14. G7- Test #091 100 50 0 100 50 100 50 100 50 0 Filt ={ 'Modulus', [50 25];... 'R21', 0.97;... 'R22', 0.97;... 'R23', 0.97;... 'seg_end', 950;... % 'AAR2', 0.25;... 'MAR1', 0.012;... % 'MAR2', 1.4;... 'MAR4', 0.032;... % 'AAR4', 0.02;... 'P*', 0.09;... 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dH', 0;... % 'h_change', [0.35 0.18];... 'dP', 0.0004};... 30 35 40 45 Modulus [GPa] 100 50 0 0.97 0.98 0.99 1 1.01 R2 100 50 0 150 200 250 300 Modulus Length [#points] 100 50 0 -1.4 -1.2 -1 -0.8 -0.6 H r [nm] 2.8 3 3.2 3.4 -3 x 10 0 Fit1 Avg.Abs.Res. 200 100 0 0.008 0.009 0.01 0.011 0.012 Fit1 Max.Abs.Res. 50 0 0.19 0.2 0.21 0.22 Fit2 Avg.Abs.Res. 0 0.5 1 1.5 50 0 Fit2 Max.Abs.Res. 0 0.1 0.2 0.3 0.4 0 h change [nm/nm] 0 0.1 0.2 0.3 0.4 100 50 0 P change [mN/mN] 100 50 0 0.008 0.009 0.01 0.011 0.012 Fit4 Avg.Abs.Res. 50 0 0.026 0.028 0.03 0.032 Fit4 Max.Abs.Res. 0.06 0.07 0.08 0.09 P* [mN] 9 10 11 12 100 50 0 h* [nm] 0 1 2 3 4 -4 x 10 40 20 0 dP [mN] 0 0.2 0.4 0.6 0.8 100 50 0 dH [nm] effective sample Fit1 Fit2 Fit3
- 15. G50- Test #185 Estat = mean: 41.7459 median: 40.9991 stdev: 2.3900 min: 38.4868 max: 48.7338 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=38.0067; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 1 0 -1 -2 -3 -4 -5 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 16. G7- Test #185 200 100 0 35 40 45 50 Modulus [GPa] 500 0 0.98 0.985 0.99 0.995 1 R2 200 100 0 150 200 250 300 350 Modulus Length [#points] 200 100 0 -1.2 -1 -0.8 -0.6 -0.4 H r [nm] 2 3 4 5 -3 x 10 200 100 0 Fit1 Avg.Abs.Res. 200 100 0 0.005 0.01 0.015 0.02 Fit1 Max.Abs.Res. 200 100 0 0.17 0.175 0.18 0.185 0.19 Fit2 Avg.Abs.Res. 200 100 0 0.5 0.6 0.7 0.8 0.9 Fit2 Max.Abs.Res. 0 0.1 0.2 0.3 0.4 400 200 0 h change [nm/nm] 0 0.05 0.1 0.15 0.2 200 100 0 P change [mN/mN] 200 100 0 0.01 0.012 0.014 0.016 Fit4 Avg.Abs.Res. 0.04 0.045 0.05 200 100 0 Fit4 Max.Abs.Res. 200 100 0 0.04 0.06 0.08 0.1 P* [mN] 9 10 11 12 400 200 0 h* [nm] 0 1 2 3 4 -4 x 10 200 100 0 dP [mN] 400 200 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 800;... 'AAR2', 0.188;... % 'MAR1', 0.015;... 'MAR2', 0.8;... 'MAR4', 0.046;... % 'AAR4', 0.02;... % 'P*', 0.05;... % 'ModLength', 300;}... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.0005};...
- 17. G7- Test #121 Estat = mean: 37.3867 median: 36.9801 stdev: 2.0155 min: 34.3962 max: 44.2580 0 0.02 0.04 0.06 0.08 0.1 0.12 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=33.9627; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 2 0 -2 -4 -6 -8 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 0.1 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 18. G7- Test #121 1000 500 0 1000 500 1000 500 0 1000 500 0 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... % 'seg_end', 800;... 'AAR2', 0.15;... 'MAR1', 0.015;... 'MAR2', 0.8;... 'MAR4', 0.045;... % % 'AAR4', 0.02;... 'P*', 0.06;... 'ModLength', 200;... % % 'h*', 95.5;... % % 'AAR2', 0.22;... 'dP', 0.0003};... 30 35 40 45 Modulus [GPa] 2000 1000 0 0.98 0.99 1 1.01 R2 500 0 200 300 400 500 Modulus Length [#points] 1000 500 0 -1 -0.8 -0.6 -0.4 -0.2 H r [nm] 1.5 2 2.5 3 3.5 -3 x 10 0 Fit1 Avg.Abs.Res. 1000 500 0 0.005 0.01 0.015 Fit1 Max.Abs.Res. 1000 500 0 0.11 0.12 0.13 0.14 0.15 Fit2 Avg.Abs.Res. 1000 500 0 0.4 0.5 0.6 0.7 0.8 Fit2 Max.Abs.Res. -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 6 8 10 12 -3 x 10 1000 500 0 Fit4 Avg.Abs.Res. 1000 500 0 0.025 0.03 0.035 0.04 0.045 Fit4 Max.Abs.Res. 0.03 0.04 0.05 0.06 P* [mN] 1000 500 0 6.5 7 7.5 8 8.5 h* [nm] 0 1 2 3 4 -4 x 10 500 0 dP [mN] 2000 1000 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3
- 19. G7- Test #142 Estat = mean: 39.9957 median: 39.7514 stdev: 2.2338 min: 35.1014 max: 45.3373 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=36.3708; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 2 0 -2 -4 -6 -8 -10 -12 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 20. G7- Test #142 1000 500 0 30 35 40 45 50 Modulus [GPa] 2000 1000 0 0.985 0.99 0.995 1 1.005 R2 1000 500 0 200 300 400 500 Modulus Length [#points] 1000 500 0 -1.6 -1.4 -1.2 -1 -0.8 H r [nm] 3 4 5 6 7 -3 x 10 2000 1000 0 Fit1 Avg.Abs.Res. 1000 500 0 0.01 0.02 0.03 0.04 Fit1 Max.Abs.Res. 1000 500 0 0.2 0.25 0.3 0.35 0.4 Fit2 Avg.Abs.Res. 1000 500 0 0.5 1 1.5 2 Fit2 Max.Abs.Res. 1000 500 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 1000 500 0 0.008 0.01 0.012 0.014 0.016 Fit4 Avg.Abs.Res. 2000 1000 0 0.02 0.03 0.04 0.05 Fit4 Max.Abs.Res. 0 0.05 0.1 0.15 0.2 1000 500 0 P* [mN] 1000 500 0 10 12 14 16 h* [nm] 0 2 4 6 -4 x 10 1000 500 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 1 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.985;... 'R22', 0.985;... 'R23', 0.985;... 'seg_end', 935;... 'AAR2', 0.4;... % % 'MAR1', 0.015;... % 'MAR2', 1.4;... % 'MAR4', 0.05;... % % % % 'AAR4', 0.02;... 'P*', 0.15;... % % 'ModLength', 200;... % % % % 'h*', 95.5;... % % % % 'AAR2', 0.22;... 'dP', 0.0006};...
- 21. G7- Test #147 Estat = mean: 36.1313 median: 36.0800 stdev: 0.7483 min: 34.6172 max: 37.9730 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.1 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=32.7217; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 14 12 10 8 6 4 2 0 -200 0 200 400 600 800 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 1 0 -1 -2 -3 -4 -5 5 -2 0 2 4 6 4 x 10 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 0.1 3500 3000 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 22. G7- Test #147 500 0 30 32 34 36 38 Modulus [GPa] 2000 1000 0 0.98 0.99 1 1.01 R2 500 0 200 300 400 500 Modulus Length [#points] 500 0 -1 -0.8 -0.6 -0.4 -0.2 H r [nm] 1 2 3 4 5 -3 x 10 1000 500 0 Fit1 Avg.Abs.Res. 0 0.01 0.02 0.03 2000 1000 0 Fit1 Max.Abs.Res. 0.16 0.18 0.2 500 0 Fit2 Avg.Abs.Res. 0 0.5 1 1.5 500 0 Fit2 Max.Abs.Res. 0 0.1 0.2 0.3 0.4 1000 500 0 h change [nm/nm] 0 0.05 0.1 0.15 0.2 500 0 P change [mN/mN] 4 6 8 10 -3 x 10 400 200 0 Fit4 Avg.Abs.Res. 1000 500 0 0.02 0.025 0.03 0.035 0.04 Fit4 Max.Abs.Res. 500 0 0.03 0.04 0.05 0.06 P* [mN] 5 5.5 6 6.5 7 500 0 h* [nm] 0 1 2 3 4 -4 x 10 400 200 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... % 'seg_end', 935;... % 'AAR2', 0.4;... % % % 'MAR1', 0.015;... % % 'MAR2', 1.4;... % % 'MAR4', 0.05;... % % % % % 'AAR4', 0.02;... 'P*', 0.05;... % % % 'ModLength', 200;... % % % % % 'h*', 95.5;... % % % % % 'AAR2', 0.22;... 'dP', 0.00025};...
- 23. G7- Test #145 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=35.9082; Bunge=0, 0, 0; Strength=0.18621; Hardening=11.3102 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data Yield Point 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 0 -1 -2 -3 -4 -5 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic Estat = mean: 39.5085 median: 39.0933 stdev: 1.4432 min: 36.7875 max: 44.3935
- 24. G7- Test #145 400 200 0 30 35 40 45 Modulus [GPa] 200 100 0 0.98 0.99 1 1.01 R2 200 100 0 150 200 250 300 350 Modulus Length [#points] 400 200 0 -1.4 -1.2 -1 -0.8 -0.6 H r [nm] 3 3.5 4 4.5 5 -3 x 10 400 200 0 Fit1 Avg.Abs.Res. 500 0 0.01 0.015 0.02 0.025 Fit1 Max.Abs.Res. 200 100 0 0.12 0.125 0.13 0.135 0.14 Fit2 Avg.Abs.Res. 400 200 0 0.4 0.45 0.5 Fit2 Max.Abs.Res. 0.4 0.5 0.6 0.7 200 100 0 h change [nm/nm] 200 100 0 0.1 0.2 0.3 0.4 0.5 P change [mN/mN] 200 100 0 0.008 0.01 0.012 0.014 Fit4 Avg.Abs.Res. 400 200 0 0.03 0.04 0.05 0.06 Fit4 Max.Abs.Res. 200 100 0 0.045 0.05 0.055 0.06 P* [mN] 200 100 0 12 12.5 13 13.5 h* [nm] 0 1 2 3 4 -4 x 10 100 50 0 dP [mN] 200 100 0 -1 -0.5 0 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... % 'seg_end', 935;... 'AAR2', 0.138;... % % % 'MAR1', 0.015;... 'MAR2', 0.5;... 'MAR4', 0.05;... % % % % % 'AAR4', 0.02;... 'P*', 0.06;... % % % % 'ModLength', 200;... % % % % % % 'h*', 95.5;... % % % % % % 'AAR2', 0.22;... 'dP', 0.0003};...
- 25. G50- Test #199 Estat = mean: 44.6301 median: 44.6072 stdev: 2.5686 min: 38.9757 max: 49.9887 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.1 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=40.7666; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 0 -5 -10 -15 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 0.1 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 26. G50- Test #199 200 100 0 35 40 45 50 Modulus [GPa] 1000 500 0 0.98 0.99 1 1.01 R2 400 200 0 150 200 250 300 350 Modulus Length [#points] 400 200 0 -1.6 -1.4 -1.2 -1 -0.8 H r [nm] 4 5 6 7 -3 x 10 500 0 Fit1 Avg.Abs.Res. 400 200 0 0.01 0.015 0.02 0.025 0.03 Fit1 Max.Abs.Res. 400 200 0 0.2 0.25 0.3 Fit2 Avg.Abs.Res. 0.8 1 1.2 1.4 200 100 0 Fit2 Max.Abs.Res. 400 200 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 400 200 0 P change [mN/mN] 400 200 0 0.008 0.01 0.012 0.014 0.016 Fit4 Avg.Abs.Res. 400 200 0 0.02 0.03 0.04 0.05 Fit4 Max.Abs.Res. 200 100 0 0.08 0.1 0.12 0.14 0.16 P* [mN] 200 100 0 28 29 30 31 32 h* [nm] 0 2 4 6 -4 x 10 200 100 0 dP [mN] 400 200 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 900;... 'AAR2', 0.5;... 'MAR1', 0.028;... 'MAR2', 1.1;... 'MAR4', 0.05;... % 'AAR4', 0.02;... 'P*', 0.4;... 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.0006};...
- 27. G7- Test #043 Estat = mean: 42.1256 median: 42.0918 stdev: 1.3368 min: 39.0036 max: 45.8276 1.4 1.2 1 0.8 0.6 0.4 0.2 0 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 Ind. Strain Ind. Stress [GPa] Es=38.328; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 12 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 2 0 -2 -4 -6 -8 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 20 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 28. G7- Test #043 400 200 0 35 40 45 50 Modulus [GPa] 1000 500 0 0.98 0.985 0.99 0.995 1 R2 500 0 250 300 350 400 450 Modulus Length [#points] 500 0 -1.4 -1.2 -1 -0.8 -0.6 H r [nm] 3 3.5 4 4.5 5 -3 x 10 1000 500 0 Fit1 Avg.Abs.Res. 1000 500 0 0.01 0.015 0.02 0.025 0.03 Fit1 Max.Abs.Res. 400 200 0 0.18 0.2 0.22 0.24 Fit2 Avg.Abs.Res. 500 0 0.5 1 1.5 Fit2 Max.Abs.Res. 1000 500 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.05 0.1 0.15 0.2 1000 500 0 P change [mN/mN] 400 200 0 0.009 0.01 0.011 0.012 0.013 Fit4 Avg.Abs.Res. 1000 500 0 0.025 0.03 0.035 0.04 0.045 Fit4 Max.Abs.Res. 400 200 0 0.02 0.04 0.06 0.08 P* [mN] 9 10 11 12 500 0 h* [nm] 0 1 2 3 4 -4 x 10 400 200 0 dP [mN] 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 850;... 'AAR2', 0.23;... % 'MAR1', 0.028;... 'MAR2', 1.25;... 'MAR4', 0.042;... % % 'AAR4', 0.02;... 'P*', 0.5;... 'ModLength', 100;... % % 'h*', 95.5;... % % 'AAR2', 0.22;... 'dP', 0.0006};...
- 29. G7- Test #049 Estat = mean: 37.9165 median: 37.7540 stdev: 1.8623 min: 34.1689 max: 42.8770 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.1 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=34.403; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 1000 2000 3000 4000 10 8 6 4 2 0 -2 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 7 0 1 2 3 4 4 x 10 0 -2 -4 -6 -8 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 30. G7- Test #049 200 100 0 30 35 40 45 Modulus [GPa] 400 200 0 0.985 0.99 0.995 1 1.005 R2 200 100 0 200 250 300 350 400 Modulus Length [#points] 200 100 0 -1 -0.9 -0.8 -0.7 -0.6 H r [nm] 2 3 4 5 -3 x 10 400 200 0 Fit1 Avg.Abs.Res. 500 0 0.01 0.02 0.03 0.04 0.05 Fit1 Max.Abs.Res. 0.15 0.16 0.17 200 100 0 Fit2 Avg.Abs.Res. 200 100 0 0.4 0.6 0.8 1 Fit2 Max.Abs.Res. 0 0.5 1 400 200 0 h change [nm/nm] 200 100 0 0.1 0.15 0.2 P change [mN/mN] 200 100 0 0.008 0.01 0.012 0.014 Fit4 Avg.Abs.Res. 1000 500 0 0.02 0.03 0.04 0.05 0.06 Fit4 Max.Abs.Res. 200 100 0 0.03 0.04 0.05 0.06 P* [mN] 200 100 0 3290.5 3291 3291.5 3292 3292.5 h* [nm] 0 1 2 3 4 -4 x 10 100 50 0 dP [mN] 200 100 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.985;... 'R22', 0.985;... 'R23', 0.985;... 'seg_end', 780;... 'AAR2', 0.22;... 'MAR1', 0.05;... 'MAR2', 2;... 'MAR4', 0.057;... 'AAR4', 0.02;... 'P*', 0.3;... % 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.00025};...
- 31. G7- Test #061 Estat = mean: 43.0638 median: 42.9278 stdev: 1.1858 min: 40.1472 max: 46.2930 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 1.2 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=39.2765; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 1 0 -1 -2 -3 -4 -5 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 32. G7- Test #061 500 0 2000 1000 1000 500 0 1000 500 0 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 750;... 'AAR2', 0.25;... % 'MAR1', 0.05;... 'MAR2', 1.2;... 'MAR4', 0.045;... % 'AAR4', 0.02;... 'P*', 0.043;... % % 'ModLength', 100;... % % 'h*', 95.5;... % % 'AAR2', 0.22;... 'dP', 0.00022};... 35 40 45 50 Modulus [GPa] 1000 500 0 0.98 0.99 1 1.01 R2 400 200 0 250 300 350 400 Modulus Length [#points] 1000 500 0 -0.7 -0.6 -0.5 -0.4 -0.3 H r [nm] 2 2.5 3 3.5 4 -3 x 10 0 Fit1 Avg.Abs.Res. 0 0.01 0.02 0.03 4000 2000 0 Fit1 Max.Abs.Res. 0.16 0.18 0.2 2000 1000 0 Fit2 Avg.Abs.Res. 500 0 0.4 0.6 0.8 1 Fit2 Max.Abs.Res. -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 400 200 0 P change [mN/mN] 8 9 10 11 -3 x 10 500 0 Fit4 Avg.Abs.Res. 1000 500 0 0.025 0.03 0.035 0.04 0.045 Fit4 Max.Abs.Res. 0.03 0.035 0.04 0.045 P* [mN] 1000 500 0 8.5 9 9.5 10 h* [nm] 0 1 2 3 4 -4 x 10 400 200 0 dP [mN] 1000 500 0 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3
- 33. G7- Test #073 Estat = mean: 40.6975 median: 39.8641 stdev: 2.5857 min: 36.6567 max: 48.4627 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 1.2 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=36.9948; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 0 -1 -2 -3 -4 -5 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 34. G7- Test #073 1000 500 0 1000 500 1000 500 0 1000 500 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 750;... 'AAR2', 0.15;... % % 'MAR1', 0.05;... 'MAR2', 0.79;... % 'MAR4', 0.045;... 'AAR4', 0.01;... 'P*', 0.06;... % % % 'ModLength', 100;... % % % 'h*', 95.5;... % % % 'AAR2', 0.22;... 'dP', 0.0003};... 30 35 40 45 50 Modulus [GPa] 2000 1000 0 0.98 0.985 0.99 0.995 1 1.005 R2 500 0 100 200 300 400 500 Modulus Length [#points] 500 0 -0.9 -0.8 -0.7 -0.6 -0.5 -0.4 H r [nm] 1.5 2 2.5 3 3.5 -3 x 10 0 Fit1 Avg.Abs.Res. 1000 500 0 0.005 0.01 0.015 0.02 Fit1 Max.Abs.Res. 2000 1000 0 0.1 0.12 0.14 0.16 Fit2 Avg.Abs.Res. 0.4 0.5 0.6 0.7 0.8 1000 500 0 Fit2 Max.Abs.Res. -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 7 8 9 10 -3 x 10 1000 500 0 Fit4 Avg.Abs.Res. 4000 2000 0 0.02 0.03 0.04 0.05 0.06 0.07 Fit4 Max.Abs.Res. 0.04 0.045 0.05 0.055 0.06 0 P* [mN] 1000 500 0 12 12.5 13 13.5 14 h* [nm] 0 1 2 3 4 -4 x 10 500 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3
- 35. G7- Test #096 Estat = mean: 46.0647 median: 46.1368 stdev: 2.4417 min: 41.0361 max: 49.9988 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=42.1943; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 0 -2 -4 -6 -8 -10 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 36. G7- Test #096 40 20 0 35 40 45 50 Modulus [GPa] 200 100 0 0.98 0.985 0.99 0.995 1 R2 100 50 0 100 150 200 250 Modulus Length [#points] 50 0 -1.4 -1.2 -1 -0.8 -0.6 H r [nm] 3 3.5 4 4.5 5 -3 x 10 100 50 0 Fit1 Avg.Abs.Res. 200 100 0 0.005 0.01 0.015 0.02 Fit1 Max.Abs.Res. 50 0 0.18 0.2 0.22 0.24 Fit2 Avg.Abs.Res. 0 0.5 1 1.5 50 0 Fit2 Max.Abs.Res. 100 50 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 200 100 0 P change [mN/mN] 100 50 0 0.008 0.009 0.01 0.011 0.012 Fit4 Avg.Abs.Res. 100 50 0 0.024 0.026 0.028 0.03 Fit4 Max.Abs.Res. 0.16 0.18 0.2 40 20 0 P* [mN] 40 20 0 19 20 21 22 h* [nm] 0 1 2 3 4 -4 x 10 40 20 0 dP [mN] 50 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 950;... 'AAR2', 0.22;... % 'MAR1', 0.05;... 'MAR2', 1.2;... 'MAR4', 0.03;... 'AAR4', 0.015;... 'P*', 0.3;... % 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.0003};...
- 37. G7- Test #097 Estat = mean: 38.6250 median: 38.2067 stdev: 2.2352 min: 35.1277 max: 44.8539 0 0.02 0.04 0.06 0.08 0.1 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=35.0619; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 0 100 200 300 400 500 10 8 6 4 2 0 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 0 -2 -4 -6 -8 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 0.1 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 38. 1000 500 G7- Test #097 0 30 35 40 45 Modulus [GPa] 4000 2000 0 0.98 0.99 1 1.01 R2 1000 500 0 100 200 300 400 500 Modulus Length [#points] 1000 500 0 -1 -0.8 -0.6 -0.4 -0.2 H r [nm] 2000 1000 1.5 2 2.5 3 3.5 -3 x 10 0 Fit1 Avg.Abs.Res. 2000 1000 0 0.005 0.01 0.015 Fit1 Max.Abs.Res. 1000 500 0 0.1 0.12 0.14 0.16 0.18 Fit2 Avg.Abs.Res. 1000 500 0 0.4 0.6 0.8 1 Fit2 Max.Abs.Res. 2000 1000 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 4 6 8 10 -3 x 10 1000 500 0 Fit4 Avg.Abs.Res. 1000 500 0 0.025 0.03 0.035 0.04 0.045 Fit4 Max.Abs.Res. 1000 500 0 0.03 0.04 0.05 0.06 P* [mN] 1000 500 0 12.5 13 13.5 14 14.5 h* [nm] 0 1 2 3 4 -4 x 10 1000 500 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... 'seg_end', 900;... 'AAR2', 0.18;... 'MAR1', 0.015;... 'MAR2', 0.9;... 'MAR4', 0.042;... 'AAR4', 0.01;... 'P*', 0.06;... % % 'ModLength', 100;... % % 'h*', 95.5;... % % 'AAR2', 0.22;... 'dP', 0.0003};...
- 39. G7- Test #108
- 40. G7- Test #108
- 41. G7- Test #131 Estat = mean: 32.5654 median: 32.3247 stdev: 2.2768 min: 28.1272 max: 40.2771 0.6 0.5 0.4 0.3 0.2 0.1 0 0 0.01 0.02 0.03 0.04 0.05 0.06 Ind. Strain Ind. Stress [GPa] Es=29.1482; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 1 0 -1 -2 -3 -4 -5 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 3000 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 42. G7- Test #131 1000 500 0 25 30 35 40 Modulus [GPa] 2000 1000 0 0.94 0.96 0.98 1 R2 1000 500 0 100 200 300 400 Modulus Length [#points] 1000 500 0 -1.5 -1 -0.5 0 H r [nm] 1 2 3 4 5 -3 x 10 2000 1000 0 Fit1 Avg.Abs.Res. 2000 1000 0 0.005 0.01 0.015 0.02 0.025 Fit1 Max.Abs.Res. 2000 1000 0 0.16 0.18 0.2 0.22 Fit2 Avg.Abs.Res. 1000 500 0 0.5 1 1.5 Fit2 Max.Abs.Res. 2000 1000 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.05 0.1 0.15 0.2 2000 1000 0 P change [mN/mN] 2000 1000 0 0.008 0.01 0.012 0.014 Fit4 Avg.Abs.Res. 2000 1000 0 0.025 0.03 0.035 0.04 Fit4 Max.Abs.Res. 1000 500 0 0.02 0.04 0.06 0.08 P* [mN] 7 8 9 10 11 1000 500 0 h* [nm] 0 1 2 3 4 -4 x 10 1000 500 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 1 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.95;... 'R22', 0.95;... 'R23', 0.95;... 'seg_end', 800;... 'AAR2', 0.22;... % 'MAR1', 0.02;... 'MAR2', 1.3;... 'MAR4', 0.04;... % % 'AAR4', 0.01;... 'P*', 0.4;... 'ModLength', 100;... % % % % 'h*', 95.5;... % % % % 'AAR2', 0.22;... 'dP', 0.0003};...
- 43. G7- Test #133 Estat = mean: 43.9060 median: 43.5871 stdev: 2.2103 min: 40.4404 max: 50.0000 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 1.4 1.2 1 0.8 0.6 0.4 0.2 0 Ind. Strain Ind. Stress [GPa] Es=40.0529; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 12 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 1 0 -1 -2 -3 -4 -5 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 44. G7- Test #133 1000 500 0 35 40 45 50 Modulus [GPa] 4000 2000 0 0.97 0.98 0.99 1 1.01 R2 1000 500 0 200 300 400 500 Modulus Length [#points] 1000 500 0 -1.5 -1 -0.5 0 H r [nm] 2 4 6 8 -3 x 10 2000 1000 0 Fit1 Avg.Abs.Res. 4000 2000 0 0.01 0.02 0.03 0.04 Fit1 Max.Abs.Res. 0.2 0.25 0.3 0.35 1000 500 0 Fit2 Avg.Abs.Res. 2000 1000 0 0.5 1 1.5 Fit2 Max.Abs.Res. 2000 1000 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 2000 1000 0 P change [mN/mN] 1000 500 0 0.008 0.01 0.012 0.014 0.016 Fit4 Avg.Abs.Res. 2000 1000 0 0.02 0.04 0.06 0.08 Fit4 Max.Abs.Res. 2000 1000 0 0.02 0.04 0.06 0.08 0.1 P* [mN] 8 9 10 11 12 2000 1000 0 h* [nm] 0 1 2 3 4 -4 x 10 1000 500 0 dP [mN] 2000 1000 0 -1 -0.5 0 0.5 1 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.97;... 'R22', 0.97;... 'R23', 0.97;... 'seg_end', 920;... 'AAR2', 0.32;... 'MAR1', 0.04;... 'MAR2', 1.5;... 'MAR4', 0.08;... % 'AAR4', 0.01;... 'P*', 0.1;... % 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.0003};...
- 45. G7- Test #157 Estat = mean: 43.9443 median: 43.6862 stdev: 2.5625 min: 39.0347 max: 49.9998 1 0.8 0.6 0.4 0.2 0 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 Ind. Strain Ind. Stress [GPa] Es=40.0814; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 12 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 4 x 10 1 0 -1 -2 -3 -4 -5 -6 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.1 0.2 0.3 0.4 0.5 12 10 8 6 4 2 0 -2 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 0.08 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 46. 400 200 G7- Test #157 0 35 40 45 50 Modulus [GPa] 1000 500 0 0.985 0.99 0.995 1 1.005 R2 400 200 0 100 200 300 400 Modulus Length [#points] 400 200 0 -1 -0.8 -0.6 -0.4 -0.2 H r [nm] 1000 500 1.5 2 2.5 3 3.5 -3 x 10 0 Fit1 Avg.Abs.Res. 6 8 10 12 -3 x 10 1000 500 0 Fit1 Max.Abs.Res. 400 200 0 0.11 0.12 0.13 0.14 0.15 Fit2 Avg.Abs.Res. 500 0 0.2 0.4 0.6 0.8 1 Fit2 Max.Abs.Res. 500 0 -0.2 0 0.2 0.4 0.6 h change [nm/nm] 0 0.1 0.2 0.3 0.4 500 0 P change [mN/mN] 6 7 8 9 10 -3 x 10 400 200 0 Fit4 Avg.Abs.Res. 1000 500 0 0.02 0.03 0.04 0.05 Fit4 Max.Abs.Res. 400 200 0 0.05 0.06 0.07 0.08 P* [mN] 400 200 0 9.5 10 10.5 11 h* [nm] 0 1 2 3 4 -4 x 10 300 200 100 dP [mN] 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [50 25];... 'R21', 0.985;... 'R22', 0.985;... 'R23', 0.985;... 'seg_end', 780;... 'AAR2', 0.138;... % % 'MAR1', 0.04;... 'MAR2', 1;... 'MAR4', 0.05;... 'AAR4', 0.01;... 'P*', 0.07;... % % % 'ModLength', 100;... % % % 'h*', 95.5;... % % % 'AAR2', 0.22;... 'dP', 0.0003};...
- 47. G7- Test #186 Estat = mean: 41.9941 median: 41.3287 stdev: 2.7468 min: 38.0638 max: 50.0335 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Ind. Strain Ind. Stress [GPa] Es=38.2265; Bunge=0, 0, 0; Strength=0; Hardening=0 Stress-Strain Modulus Line Hardening Fit Strain Offset Modulus Fit Data Hardening Fit Data 10 8 6 4 2 0 -200 0 200 400 600 displacement / nm load / mN Load Vs. Displacement Raw Data 0 Pt. Data 5 0 1 2 3 4 5 4 x 10 2 0 -2 -4 -6 -8 x 10 S P2/3-Sh Zero Point Fit Raw Data 0 Pt. Data 0 0.2 0.4 0.6 0.8 15 10 5 0 -5 P 2/3 h Modulus Fit Data Elastic 0 0.02 0.04 0.06 2500 2000 1500 1000 500 0 Strain Contact Radius / nm Strain Vs. Contact Radius Data Elastic
- 48. G7- Test #133 1000 500 0 30 35 40 45 50 Modulus [GPa] 2000 1000 0 0.98 0.985 0.99 0.995 1 R2 1000 500 0 100 200 300 400 Modulus Length [#points] 1000 500 0 -1.4 -1.2 -1 -0.8 -0.6 H r [nm] 1000 500 3 4 5 6 -3 x 10 0 Fit1 Avg.Abs.Res. 1000 500 0 0.01 0.015 0.02 0.025 Fit1 Max.Abs.Res. 2000 1000 0 0.16 0.18 0.2 0.22 Fit2 Avg.Abs.Res. 0 0.5 1 1.5 1000 500 0 Fit2 Max.Abs.Res. 0 0.2 0.4 0.6 0.8 1000 500 0 h change [nm/nm] 0 0.1 0.2 0.3 0.4 1000 500 0 P change [mN/mN] 1000 500 0 0.01 0.012 0.014 0.016 Fit4 Avg.Abs.Res. 1000 500 0 0.025 0.03 0.035 0.04 0.045 Fit4 Max.Abs.Res. 1000 500 0 0.04 0.06 0.08 0.1 0.12 P* [mN] 8 10 12 14 1000 500 0 h* [nm] 0 1 2 3 4 -4 x 10 400 200 0 dP [mN] 1000 500 0 -1 -0.5 0 0.5 dH [nm] effective sample Fit1 Fit2 Fit3 Filt ={ 'Modulus', [60 25];... 'R21', 0.98;... 'R22', 0.98;... 'R23', 0.98;... % 'seg_end', 780;... 'AAR2', 0.22;... % 'MAR1', 0.04;... 'MAR2', 1.2;... 'MAR4', 0.045;... 'AAR4', 0.016;... 'P*', 0.15;... % 'ModLength', 100;... % 'h*', 95.5;... % 'AAR2', 0.22;... 'dP', 0.0004};...
- 49. G7- Test #133
- 50. G7- Test #133