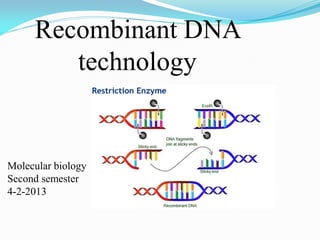
Dna recombinant technology
- 1. Recombinant DNA technology Molecular biology Second semester 4-2-2013
- 2. A series of procedures used to recombine DNA segments. Under certain conditions, a recombinant DNA molecule can enter a cell and replicate.
- 3. Recombinant DNA technology is one of the recent advances in biotechnology, which was developed by two scientists named Boyer and Cohen in 1973.
- 4. RECOMBINANT DNA RECOMBINANT DNA:- DNA molecules constructed outside of living cells by joining natural or synthetic DNA segments to DNA molecules that can replicate in a living cell
- 5. The DNA is inserted into another DNA molecule called ‘vector’ The recombinant vector is then introduced into a host cell where it replicates itself, the gene is then produced
- 6. Basic principle of recombinant DNA technology
- 7. How is Recombinant DNA made? There are three different methods by which Recombinant DNA is made. They are Transformation, Phage Introduction, and Non-Bacterial Transformation.
- 8. Transformation The first step in transformation is to select a piece of DNA to be inserted into a vector. The second step is to cut that piece of DNA with a restriction enzyme and then ligate the DNA insert into the vector with DNA Ligase. The insert contains a selectable marker which allows for identification of recombinant molecules. The vector is inserted into a host cell, in a process called transformation. One example of a possible host cell is E. coli. The host cells must be specially prepared to take up the foreign DNA.
- 9. Non-Bacterial transformation Microinjection, the DNA is injected directly into the nucleus of the cell being transformed. The host cells are bombarded with high velocity micro-projectiles, such as particles of gold or tungsten that have been coated with DNA.
- 10. Phage Introduction Phage introduction is the process of transfection, which is equivalent to transformation, except a phage is used instead of bacteria. In vitro packaging of a vector is used. This uses lambda or MI3 phages to produce phage plaques which contain recombinants. The recombinants that are created can be identified by differences in the recombinants and non-recombinants using various selection methods.
- 11. How does rDNA work? Recombinant DNA works when the host cell expresses protein from the recombinant genes. A significant amount of recombinant protein will not be produced by the host unless expression factors are added. Protein expression depends upon the gene being surrounded by a collection of signals which provide instructions for the transcription and translation of the gene by the cell. These signals include the promoter, the ribosome binding site, and the terminator.
- 12. Expression vectors, in which the foreign DNA is inserted, contain these signals. Signals are species specific. In the case of E. coli, these signals must be E. coli signals as E. coli is unlikely to understand the signals of human promoters and terminators. Problems are encountered if the gene contains introns or contains signals which act as terminators to a bacterial host. This results in premature termination, and the recombinant protein may not be processed correctly, be folded correctly, or may even be degraded.
- 13. Production of recombinant proteins in eukaryotic systems generally takes place in yeast and filamentous fungi. The uses of animal cells is difficult due to the fact that many need a solid support surface, unlike bacteria, and have complex growth needs. However, some proteins are too complex to be produced in bacterium, so eukaryotic cells must be used.
- 14. Large-scale production of human proteins by genetically engineered bacteria. Such as : insulin, Growth hormone, Interferons and Blood clotting factors (VIII & IX)
- 15. 1) Obtaining the human insulin gene Human insulin gene can be obtained by making a complementary DNA (cDNA) copy of the messenger RNA (mRNA) for human insulin.
- 16. 2)Joining the human insulin gene into a plasmid vector The bacterial plasmids and the cDNA are mixed together. The human insulin gene (cDNA) is inserted into the plasmid through complementary base pairing at sticky ends.
- 17. 3)Introducing the recombinant DNA plasmids into bacteria The bacteria E.coli is used as the host cell. If E. coli and the recombinant plasmids are mixed together in a test-tube.
- 18. 4)Selecting the bacteria which have taken up the correct piece of DNA The bacteria are spread onto nutrient agar. The agar also contains substances such as an antibiotic which allows growth of only the transformed bacteria.
- 19. Vaccine development The surface antigen of Plasmodium falciparum, one of the 4 species of malaria, has been transferred to E. coli to produce amounts large enough to develop a vaccine against this form of malaria. It works well enough for people who will visit a malarious region for a relatively short period of time.
- 20. Hemophilia A and B The genes encoding factors 8 and 9 are on the X chromosome. Like other X-linked disorders, hemophilia A and B are found almost exclusively in males because they inherit just a single X chromosome, and if the gene for factor 8 (or 9) on it is defective, they will suffer from the disease. There are many different mutant versions of the genes for factors 8 and 9. Although some produce only a minor effect on the function of their protein, others fail to produce any functioning clotting factor.
- 21. Transferring the gene for normal adult hemoglobin into marrow stem cells of an individual with sickle-cell anemia. The goal is to promote the growth of enough cells to produce enough normal hemoglobin to alleviate the symptoms of sickle-cell anemia. Gene therapy for genetic diseases
- 22. Safety Issues in relation to Recombinant DNA Technology As bacteria is commonly used in recombinant DNA work, there has always been a concern among scientists and a worry among people that there is a possibility that a clone of highly pathogenic recombinant bacteria were made by accident, then escaped from the laboratory and caused an epidemic for which no drugs were available. Recombinant DNA Advisory Committee (RAC) was established in 1974 in the United States, which responds to public concerns regarding the safety of manipulation of genetic material through the use of recombinant DNA techniques.
- 23. 2 types of control : physical containment and biological containment Effective biological safety programs were operated in a variety of laboratories, which include a set of standard practices generally used in microbiological laboratories, and special procedures, equipment and laboratory installations that provide physical barriers of varying degrees.
- 24. In considering biological containment, the vector (plasmid, organelle, or virus) for the recombinant DNA and the host (bacterial, plant, or animal cell) in which the vector is propagated in the laboratory will be considered together. (i) survival of the vector in its host outside the laboratory, and (ii) transmission of the vector from the propagation host to other non- laboratory hosts. Biological containment
- 25. It is always possible that an antibiotic-resistant plasmid could be accidentally incorporated into a dangerous pathogen with serious medical consequences.
- 27. Introduction Within living cells, the exchange of DNA sequences and genetic information can occur through a regulated series enzymatic reactions involving pairing of DNA molecules and phosphodiester bond breakage and rejoining. This type of sequence rearrangement is known as genetic recombination. genetic recombination responsible for rearranging sequences between different pieces of DNA, shaping the genome by altering the sequences that are present, pairing chromosome before cell division and promoting DNA repair.
- 28. DNA Recombination Roles Types
- 29. Biological Roles for Recombination 1. Generating new gene/allele combinations (crossing over during meiosis) 2. Generating new genes (e.g., Immuno- globulin rearrangement) 3. Integration of a specific DNA element 4. DNA repair
- 30. Practical Uses of Recombination 1. Used to map genes on chromosomes (recombination frequency proportional to distance between genes) 2. Making transgenic cells and organisms
- 31. Map of Chromosome I of Chlamydomonas reinhardtii cM = centiMorgan; unit of recombination frequency 1 cM = 1% recombination frequency
- 32. Types of Recombination 1. Homologous - occurs between sequences that are nearly identical (e.g., during meiosis) 2. Site-Specific - occurs between sequences with a limited stretch of similarity; involves specific sites 3. Transposition – DNA element moves from one site to another, usually little sequence similarity involved
- 33. Homologous recombination Is the exchange of DNA sequences between DNA molecules that contain identical or nearly identical sequences along their length. The common stretch of bases that will be recombined is known as the homology between the sequences and can be as few as 50-100bp or as much as a whole chromosome. The greater the region of homology the higher the frequency of recombination.
- 34. Requirement of homologous recombination 1- Two DNA sequences with similar or identical base pair sequences. 2- The ability to form stable hydrogen bonds between the bases on one strand of one DNA sequence and the base on the complementary strand on the other DNA sequence. 3- The proteins needed to carry out recombination.
- 35. Rec A, the most important protein in DNA recombination. 38 kDa protein that polymerizes onto SS DNA 5’-3’, Catalyzes strand exchange, has an ATPase, Also binds DS DNA, but not as strongly as SS
- 36. RecA, mechanism of action 3 steps of strand exchange: 1. Pre-synapsis: recA coats single stranded DNA (accelerated by SSB, get more relaxed structure) 2. Synapsis: alignment of complementary sequences in SS and DS DNA (paranemic or side-by-side structure) 3. Post-synapsis or strand-exchange: SS DNA replaces the same strand in the duplex to form a new DS DNA (requires ATP hydrolysis)
- 38. But what create a single strand DNA in the cell? Non enzymatic (physical agents) Enzymatic (specific enzyme responsible for creating ssDNA). This enzyme is known as RecBCD.
- 39. RecBCD : A complex enzyme RecBCD has several enzymatic function: 1. Endonuclease subunits (recBCD) that cut one DNA strand close to Chi sequence. 2. DNA helicase activity (recBC subunit) and 3. DNA-dependent ATPase activity unwinds DNA to generate SS regions
- 41. recBCD Pathway of Homologous Recombination Part I: Nicking and Exchanging 1. A nick is created in one strand by recBCD at a Chi sequence (GCTGGTGG), found every 5000 bp. 2. Unwinding of DNA containing Chi sequence by recBCD allows binding of SSB and recA. 3. recA promotes strand invasion into homologous DNA, displacing one strand. 4. The displaced strand base-pairs with the single strand left behind on the other chromosome. 5. The displaced and now paired strand is nicked (by recBCD?) to complete strand exchange.
- 42. The recBCD Pathway of Homologous Recombination Part I: Nicking and Exchanging
- 43. recBCD Pathway of Homologous Rec. Part II: Branch Migration and Resolution 1. Nicks are sealed Holliday Junction 2. Branch migration (ruvA + ruvB) 3. Resolution of Holliday Junction (ruvC)
- 44. recBCD Pathway of Homologous Recombination Part II: Branch Migration and Resolution
- 45. RuvA and RuvB DNA helicase that catalyzes branch migration RuvA tetramer binds to HJ (each DNA helix between subunits) RuvB is a hexamer ring, has helicase & ATPase activity 2 copies of ruvB bind at the HJ (to ruvA and 2 of the DNA helices) Branch migration is in the direction of recA mediated strand-exchange
- 47. RuvC : resolvase Endonuclease that cuts 2 strands of HJ Binds to HJ as a dimer. Consensus sequence: (A/T)TT (G/C) - occurs frequently in E. coli genome - branch migration needed to reach consensus sequence!
- 49. Models of recombination: 1- copy choice model: • In this model replication is a requirement for recombination (DNA pol III switches positions, alternating back and forth of between two DNA molecules acting as replication template).
- 51. 2- Holliday Model R. Holliday (1964) - Holliday Junctions form during recombination - HJs can be resolved 2 ways, only one produces true recombinant molecules
- 52. EM of a Holliday Junction w/a few melted base pairs around junction
- 53. 3
- 54. DNA exchange in bacteria Plasmids Methods of DNA 2nd semeste 20-2-2012
- 55. Introduction Bacteria can exchange or transfer DNA between other bacteria in three different ways. In every case the source cells of the DNA are called the DONORS and the cells that receive the DNA are called the RECIPIENTS. In each case the donor DNA is incorporated into the recipients cell's DNA by recombination exchange. 55
- 56. Plasmids are pieces of DNA that exist separate from the chromosome. They contain an Origin of replication (ori) and, as such Replicate independentely from the chromosome. Plasmid contain a variety other genes; antibiotic resistant gene, degradation of different carbon sources and genes involved in causing disease 56 Plasmids
- 57. DNA synthesis from the origin proceeding in one direction at a time (unidirectional DNA synthesis) or both directions (bidirectional DNA synthesis). 57
- 58. Replicon Because plasmids have a mechanism to replicate their DNA and ensure that they are stably maintained in a cell population, they are also known as replicon. 58
- 59. Plasmid copy number Low copy number plasmids (1-2 copy): P1 phage & F plasmid or pSC101 (10-15 copy) High copy number plasmids (50 copies): ColE1 (50) 59
- 60. Plasmid partitioning High copy no. plasmid do not have any mechanism of partitioning . Low copy no. plasmids must have a mechanism to ensure their proper partitioning. 60
- 61. parS : A specific site on low copy number plasmids required for segregation. ParA & parB proteins bind to parS sequence, 61
- 62. How parS, parA and parB protein complex function? Keep the plasmid molecule in the middle of the cell until daughter cell are clearly distinguished. How the plasmids distinguished between the different physical locations in the cell? 62
- 63. 63
- 64. Plasmid incompatibility Maintenance of more than one plasmids in the cell can only occur if the plasmids can only carry different origins of replication. The inability of two plasmids with the same origins to be maintained in the same cell is known as incompatibility. 64
- 65. 65
- 66. Incompatibility can be explained by a limiting concentration of initiator protein and random replication of plasmid molecules. Plasmid molecules are randomly chosen from the population and undergo DNA replication each time the plasmid is replicated. Different plasmids that use the same or very similar origins and initiator proteins are rapidly separated from each other. 66
- 67. Broad host range plasmids Plasmid that can be replicated and maintained in many different bacterial species are known as broad host range plasmids. 67
- 68. Moving plasmids from one cell to another. By * Conjugation (Shuttle vector) Risk: Moving antibiotics resistance between bacterial species. 68
- 69. Gene exchangeRequirement Contact Genes transferred in most Transformation Free donor DNA fragment Competent recipient cell no Most capsule gene, unlimited Conjugation Sex pili on donor Fertility plasmid in donor Live cells Closely related species yes Drug resistance, resistant genes to toxin, enzymes etc. Transduction Donor lysed by phage Defective phage carries donor DNA Live competent recipient cell no Toxins, drug resistance
- 70. Bacterial conjugation is the transfer of genetic material between bacteria through direct cell to cell contact, or through a bridge-like connection between the two cells. conjugation is a mechanism of horizontal gene transfer.
- 71. The prototype for conjugative plasmids is the F- plasmid, also called the F-factor. The F-plasmid is an episome (a plasmid that can integrate itself into the bacterial chromosome by homologous recombination) of about 100 kb length. It carries its own origin of replication, the oriV, as well as an origin of transfer, or oriT. There can only be one copy of the F-plasmid in a given bacterium, either free or integrated. The host bacterium is called F-positive or F-plus (denoted F+). Strains that lack F plasmids are called F- negative or F-minus (F-).
- 72. Among other genetic information, the F-plasmid carries a tra and a trb locus, which together are about 33 kb long and consist of about 40 genes. The tra locus includes the pilin gene and regulatory genes, which together form pili on the cell surface, polymeric proteins that can attach themselves to the surface of F- bacteria and initiate the conjugation.
- 73. F factor: the plasmid
- 74. When conjugation is initiated, a relaxase enzyme creates a nick in one plasmid DNA strand at the origin of transfer, or oriT. The relaxase may work alone or in a complex of over a dozen proteins, known collectively as a relaxosome. In the F-plasmid system, the relaxase enzyme is called TraI and the relaxosome consists of TraI, TraY, TraM, and the integrated host factor, IHF.
- 75. The transferred, or T-strand, is unwound from the duplex plasmid and transferred into the recipient bacterium in a 5'-terminus to 3'-terminus direction. The remaining strand is replicated, either independent of conjugative action beginning at the oriV) or in concert with conjugation (conjugative replication similar to the rolling circle replication of lambda phage).
- 77. If the F-plasmid becomes integrated into the host genome, donor chromosomal DNA may be transferred along with plasmid DNA and the cell is called Hfr. The certain amount of chromosomal DNA that is transferred depends on how long the bacteria remain in contact; for common laboratory strains of E. coli the transfer of the entire bacterial chromosome takes about 100 minutes. The transferred DNA can be integrated into the recipient genome via homologous recombination.
- 78. Some strains of bacteria with an integrated F-plasmid can be isolated and grown in pure culture. Because such strains transfer chromosomal genes very efficiently, they are called Hfr (high frequency of recombination). 78
- 79. Formation of the Hfr F plasmid recombine to the chromosome by homologous recombination. F plasmid carry two copies of IS3, one copy of IS2 and one copy of Tn1000. The chromosome has about six copies of IS2, five copies of IS3 and multiple copies of Tn1000. These sites can be used as a source of homologous recombination. 79
- 80. Formation of the F´( F prime) F´ are formed from Hfr strains. The F plasmid that is integrated into the chromosome can come back. The F factor can come back out with only F plasmid reforming F+. The F factor can come back out and carry chromosomal DNA either from left or right of the original insertion these are known as type I F prime. 80
- 81. If the F´ carry genes that were transferred early by the starting Hfr strain then it’s a Type IA F´, If the F´ carry genes that were transferred late by the starting Hfr strain then it’s a Type IB F´, Its possible for the F factor to carry genes from both sides of the original chromosome and these are called Type IIF´ 81
- 82. Genetic uses of F´ Merodiploid ( two copies of the gene into the same cell) 82
- 83. Genetic uses of the Hfr Mapping genes 83 A B C D
- 84. Lec 4 Transformation & Transduction
- 85. Transformation: Is the process that allows bacteria to take up free or naked DNA from their surrounding environment. Bacteria that have undergoes transformation are called transformants. Transformation requires that the bacterium can become naturally competent to take up DNA from their surrounding environment.
- 86. Natural competency When do bacterial cell become competent to take up DNA ? the physiological state of the cell influences its ability to become competent.
- 87. Establishment of the competent cell Based on Cell density (end of the exponential phase) Depletion carbon, nitrogen and phosphorus Transformation Sporulation
- 88. Two pathways activate the competence genes in B. subtilis. P1 sense the cell density, and when enough cells are present, signals the cells to induce the com genes. P2 monitors the available nutrients and when they become limiting, signals the cells to induce the com genes.
- 89. One pathway used by B. subtillis relies on a small peptide, com X to sense the cell density and induce the com genes. ComX is produced in the cytoplasm as a larger precursor molecule, the comQ protein processes and modifies precursor comX to make a smaller active peptide. The active comx is excreted from the cell.
- 90. Two component signal transduction systems use a membrane protein called sensor-kinase to sense an environmental signal (a). The sensor - kinase autophosphorylates on a specific histidine residue in the protein(b). The sensor- kinase transfer this phosphate to the response regulator (c). This activates the response regulator so that it can induce and/ or repress specific genes (d). These genes allow a cell to respond to the original environmental signal in an appropriate way.
- 91. ComP and comA comprise a two components signal transduction system in B. subtillis for the induction of the competency genes. (a) ComX binds to comP. (b) This binding induce autophosphorylation of comP. (c) The phosphate is relied to comA. (d) the phosphorylated comA activates the comS genes. (e) ComS leads to the induction of the competency genes.
- 92. Quorum sensing system ComP, comA and comX depend on cell density therefore they called quorum sensing system.
- 93. The second pathway to induce the competency genes in B.subtillis uses another small peptide called CSF. CSF transported to the cell via membrane protein called spo0k .
- 94. The process of transformation Transformation in B. subtillis a- A type IV pillus binds the double stranded DNA first. b- The DNA is transferred to the comEA receptor. c- An endonuclease comI recognizes the receptor with bound DNA. d- ComI degrade one strand of the DNA. e- The single stranded DNA is transported in to the cell and protected by a single stranded DNA binding protein.
- 95. Some species that are naturally competent can only take up DNA that contains specific base sequence called USS or uptake signal sequence.
- 96. Some species used a specialized structure called a transformasome to import DNA. The DNA most contain a USS and initially is double stranded. During the process of translocation, the DNA is degraded to a single – stranded molecule.
- 97. Transformation as a genetic tool: gene mapping Mapping genes requires that a piece of chromosomal DNA from one bacterium be introduced to another bacterium of the same species. Transformation as a molecular tool: Introducing genes to many different bacteria.
- 99. Aim of this lecture To understand the method of moving DNA by phage To understand generalized and specialized transducing phage To differentiate between screening and selection. To know the benefit of transduction.
- 100. Transduction: Is the process of moving a piece of chromosome (DNA ) from one cell to another using a bacteriophage to carry DNA. First described by Zinder and Lederburg in 1952.
- 101. There are two types of transducing phage 1- Generalized: capable of moving any piece of DNA from one cell to another. 2- specialized: move the same piece of DNA every time.
- 103. P1 as a model for generalized transducing phage P1 has a double stranded DNA (90kb). P1 infect E.coli and package DNA into a phage head from a specific site on the P1 DNA known as pac site, once the DNA is inserted into the phage head, a p1tail is added to make a complete phage that is capable of infecting another E.coli cell.
- 104. Specialized transducing phage Specialized transducing phage carrying a defined region of the chromosome. Unlike generalized transduing phage specialized transducing phage carry chromsomal DNA in addition to phage DNA. Example is λ.
- 105. λ recombine with the bacterial chromosome using attB site by site specific recombination.
- 106. Identifying transdused bacteria Screening: examining each individual bacterium is known as screening. Selection: bacteria with specific characteristics or phenotype can be selected by growth or non growth on specified media. A- Lethal selection B- non lethal selection
- 107. Uses for transduction 1- Two factor crosses to determine gene linkage. 2- Mapping the orders of the gene – three factor crosses. 3- strain construction 4- Localized mutagenesis. 5- Making merodiploides using specialized transduction. 6- Moving mutations from plasmids to specialized transducing phage to the chromosome.
- 108. DNA Cloning This lecture is concerned with the details of the individual steps in the cloning process: 1- How is the DNA removed from the cell? 2- How is the DNA is cut into pieces? 3- How are the pieces of DNA put back together? 4- How we do monitor each of these steps?
- 109. DNA Cloning Isolation DNA from cells. Plasmid DNA isolation. Chromosomal DNA isolation. Cutting DNA molecules (Restriction enzymes). Joining DNA molecules. Application of DNA cloning (DNA library). DNA detection 1- Southern blotting 2- DNA amplification PCR.
- 110. Plasmid DNA isolation The first step in DNA cloning is isolating of the vector and chromosomal DNA. In the general scheme, cells containing the plasmid are grown to a high cell density, gently lysed, and the plasmid DNA is isolated and concentrated.
- 111. Cells can be lysed by different method depending on the size of plasmid molecule. Most procedures used EDTA to chelate the Mg+ ² associated the outer membrane and destabilize the outer membrane. Lysosyme is added to digest the peptidoglycan and detergent are frequently used to solubilize the membrane. RNAses are added to degrade the large amount of RNA found in the actively growing cells. (for more detail see the practical part)**
- 112. Chromosomal DNA isolation To isolate chromosomal DNA, cells are lysed in the same way as for plasmid DNA isolation. The cell lysate is extracted using phenol or otherwise treated to remove all proteins. The chromosomal DNA is very fragile and breaks easily, for this reason, the chromosomal DNA is not usually purified using columns.
- 113. Cutting DNA Molecule Once DNA has been purified, it must be cut into pieces before the chromosomal DNA and the plasmid DNA can be joined. A group of enzymes called restriction enzymes are used for cutting DNA.
- 114. Why bacteria use restriction and modification enzyme? Bacteria use restriction and modification enzymes to identify their own DNA from any foreign DNA that enter their cytoplasm. The restriction part of the system is an enzyme that recognize a specific site on DNA called restriction site and cleaves DNA by catalyzing breaks in specific phosphodiester bonds on both strands of DNA. The modification part of the system is a protein that methylates the DNA sequence so that the restriction enzyme no longer recognize the DNA sequence. Thus the bacteria can protect its own DNA from the restriction enzyme.
- 115. Types of restriction enzymes 1- Type I restriction–modification systems Three different proteins form a complex that carries out both restriction and modification of the DNA. The complex must interact with a cofactor, S- adenosylmethionine, before it is capable of recognizing DNA. The S-adenosylmethionine is the methyl donor for the modification reaction and all known Type I systems methylate adenine residues on both strands of the DNA. The restriction reaction requires ATP and Mg++ for cleavage of the DNA. The DNA sequence recognized by Type I enzymes is asymmetric. Cleavage of the DNA occurs randomly, usually no closer than 400 bp from the recognition sequence and sometimes as far away as 7000 bp.
- 116. Type II restriction–modification systems Composed of two independent proteins. One protein is responsible for modifying the DNA and one for restricting the DNA. Modification of the DNA uses S- adenosylmethionine as the methyl donor. The modifications that have been found are 5-methlycytosine, 4-methylcytosine, or 6-methlyadenosine. The DNA sequence recognized by Type II restriction enzymes is symmetric and usually palindromic. Both the cleavage of the DNA and modification of the DNA occur symmetrically on both strands of the DNA within the recognition sequence. Several thousand Type II systems have been identified. Type II restriction enzymes are the most useful for cloning because they generate DNA molecules with a specific sequence on the ends.
- 117. Type III restriction–modification systems Type III systems are composed of two different proteins in a complex. The complex is responsible for both restriction and modification. Modification requires S-adenosylmethionine, is stimulated by ATP and Mg++, and occurs as 6-methyladenine. Restriction requires Mg++ and is stimulated by ATP and S- adenosylmethionine. The recognition sites for Type III enzymes are asymmetric and 5–6 bp in length. The DNA is cleaved on the 3´side of the recognition sequence, 25–27 bp away from the recognition sequence. Type III restriction enzymes require two recognition sites in inverted orientation in order to cleave the DNA.
- 118. The sequence recognized by restriction enzymes
- 119. Type II restriction enzymes generate DNA molecules with specific sequences on both ends. These ends can be rejoined to regenerate the restriction site.
- 120. Restriction–modification as a molecular tool The cleavage is on both strands of the DNA and results in a double- stranded break. Cleavage of the DNA leaves one of three types of ends, depending upon the specific restriction enzyme. Some enzymes leave a 5´ overhang, some a 3´ overhang, and some leave blunt ends. The ends with either a 5´ or 3 ´ overhang are known as sticky ends.
- 122. Joining DNA molecules The double-stranded ends must be covalently attached. A version of this reaction is normally carried out in the cell by an enzyme known as DNA ligase. The double-stranded break formed by the restriction enzyme can be thought of as two nicks, each of which is a substrate for ligase. If a plasmid molecule that has been digested with a restriction enzyme is subsequently treated with ligase, the plasmid molecule ends can be covalently closed by ligase . Ligation is an energy-requiring reaction that occurs in three distinct steps. 1- The adenylyl group from ATP is covalently attached to ligase and inorganic phosphate is released. 2- The adenylyl group is transferred from ligase to the 5´ phosphate of the DNA in the nick. 3- The phosphodiester bond is formed when the 3 ´ OH in the nick attacks the activated 5 ´ phosphate. AMP is released in the process.
- 124. DNA Libraries & Detection of cloned DNA
- 125. Aims of this lecture 1- To describe how a library of DNA can be constructed using DNA cloning method. 2- To understand how cloned DNA can be detected by southern blotting and PCR.
- 126. Introduction In 1962 the Noble prize in medicine and physiology was awarded to Watson and Crick for the discovery of the structure of DNA. The technology developed in the 50 years since has revolutionized how biological research is conducted. The ability to manipulate genes in vitro has greatly increased not only the experiments that are know possible but also how scientists think about biological problems. Each of the techniques that will be describe in this lecture allows scientists to manipulate a novel gene in many different ways with the goal of uncovering its unique role in the cell.
- 127. DNA Libraries
- 128. DNA libraries –a collections of DNA sequences. DNA libraries, like conventional libraries, are used to collect and store information. In DNA libraries, the information is stored as a set of DNA molecules. All DNA libraries are collections of DNA fragments that represent a particular biological system of interest. The two most common uses for these DNA collections are DNA sequencing and gene cloning.
- 129. A DNA library is a collection of clones of DNA designed so that there is a high probability of finding any particular piece of the source DNA in the collection. DNA libraries can be made using highly efficient cloning vectors such as lambda phages, plasmids, cosmids, P1 phages and bacterial or yeast artificial chromosomes.
- 130. Types of DNA Libraries The genomic library contains DNA fragments representing the entire genome of an organism. The cDNA library contains only complementary DNA molecules synthesized from mRNA molecules in a cell.
- 131. Genomic Library Are made from total nuclear DNA of an organism or species. DNA is cut into clonable size pieces as randomly as possible using restriction endonuclease. Genomic libraries contain whole genomic fragments including gene exons and introns, gene promoters, intragenic DNA, centromeric DNA, origins of replication, etc
- 132. Constructing libraries of clones The library is made by inserting these millions of fragments of DNA into λ bacteriophage plasmids. This allows the genes to be grown up (cloned) in E. coli. The library can be screened for DNA fragments or Particular genes
- 134. cDNA Library The advantage of cDNA library is that it contains only the coding region of a genome.
- 136. Vectors for DNA Libraries Genomic libraries – λ-phage - 9-23 kb → convenient and easy to handle – Cosmids - 30-45 – PAC, BAC, YAC →artificial chromosomes, accommodate large fragments cDNA libraries -λ-phage - 9-23 kb → provides selection for longer cDNAs -conventional plasmids → high level of expression of proteins.
- 137. DNA detection—Southern blottingIn 1975, E.M. Southern described a technique to detect sequence homology between two molecules, without determining the exact base sequence of the molecules. The Southern blot is used to detect the presence of a particular piece of DNA in a sample. The DNA detected can be a single gene, or it can be part of a larger piece of DNA such as a viral genome.
- 138. The key to this method is hybridization. Hybridization-process of forming a double-stranded DNA molecule between a single-stranded DNA probe and a single-stranded target patient DNA.
- 139. There are 2 important features of hybridization: The reactions are specific-the probes will only bind to targets with a complementary sequence. The probe can find one molecule of target in a mixture of millions of related but non- complementary molecules.
- 140. Steps for hybridization The technique relies on fractionating the DNA on an agarose gel and denaturing the fractionated DNA in the agarose. The denatured DNA is transferred to a solid support, such as a nylon or nitrocellulose filter. A second DNA, called the probe, is labeled with a tag, denatured, and applied to the filter. Probes can be tagged with radioactivity and detected with X-ray film.
- 141. They can also be labeled with fluorescent nucleotides or enzymes such as alkaline phosphatase or horseradish peroxidase. The enzymes are then detected with special substrate molecules that change color or emit light when cleaved by the enzyme. The probe will hybridize with any DNA on the filter that has complementary base sequences. Once the excess, non- hybridized probe is washed away, the tag attached to the probe can be detected.
- 142. Definition of DNA probe : a single-stranded DNA molecule used in laboratory experiments to detect the presence of a complementary sequence among a mixture of other singled-stranded DNA molecules.
- 143. SOUTHERN BLOTTING
- 144. RNA blot : Northen Blot It can be used to determine the temporal and spatial locations of RNA expression by ―running‖ an RNA blot, often referred to as a northern blot.
- 146. specific DNA detection by hybridization
- 147. PCR Based Screening DNA Library
- 148. USES Identify mutations, deletions, and gene rearrangements. Used in diagnosis of cancer and in prenatal diagnosis of genetic diseases Leukemias. Diagnosis of HIV-1 and infectious disease.
- 149. USES Every person has repeated sequences of base pairs which are called Variable Number Tandem Repeats (VNTRs) To find a particular VNTR we use a radioactive version of the one in question. This pattern is known as a DNA fingerprint.
- 150. USES Applications of DNA fingerprinting include: Paternity and Maternity Testing Criminal Identification and Forensics Personal Identification
- 151. DNA sequencing Introduction: Knowledge of DNA sequences has become indispensable for basic biological research, other research branches utilizing DNA sequencing, and in numerous applied fields such as diagnostic, biotechnology, forensic biology and biological systematics. The advent of DNA sequencing has significantly accelerated biological research and discovery. The rapid speed of sequencing attained with modern DNA sequencing technology has been instrumental in the sequencing of the human genome, in the human genome project. Related projects, often by scientific collaboration across continents, have generated the complete DNA sequences of many animal, plant, and microbial genomes.
- 152. DNA sequencing includes several methods and technologies that are used for determining the order of the nucleotide bases—adenine, guanine, cytosine, and thymine—in a molecule of DNA.
- 153. Two similar methods used for determining the order of the nucleotide sequences: 1. Maxam and Gilbert method 2. Sanger method They depend on the production of a mixture of oligonucleotides labeled either radioactively or fluorescing, with one common end and differing in length by a single nucleotide at the other end This mixture of oligonucleotides is separated by high resolution electrophoresis on polyacrilamide gels and the position of the bands determined
- 154. The Maxam-Gilbert Technique Principle - Chemical Degradation of Purines Purines (A, G) damaged by dimethylsulfate Methylation of base Heat releases base Alkali cleaves G Dilute acid cleave A>G
- 155. Maxam-Gilbert Technique Principle Chemical Degradation of Pyrimidines Pyrimidines (C, T) are damaged by hydrazine Piperidine cleaves the backbone 2 M NaCl inhibits the reaction with T
- 156. Maxam and Gilbert Method Chemical degradation of purified fragments (chemical degradation) The single stranded DNA fragment to be sequenced is end-labeled by treatment with alkaline phosphatase to remove the 5’phosphate It is then followed by reaction with P-labeled ATP in the presence of polynucleotide kinase, which attaches P labeled to the 5’terminal The labeled DNA fragment is then divided into four aliquots, each of which is treated with a reagent which modifies a specific base 1. Aliquot A + dimethyl sulphate, which methylates guanine residue 2. Aliquot B + formic acid, which modifies adenine and guanine residues 3. Aliquot C + Hydrazine, which modifies thymine + cytosine residues 4. Aliquot D + Hydrazine + 5 mol/l NaCl, which makes the reaction specific for cytosine The four are incubated with piperidine which cleaves the sugar phosphate backbone of DNA next to the residue that has been modified
- 158. Chain-termination methods (Sanger method) The key principle of the Sanger method was the use of dideoxynucleotide triphosphates (ddNTPs) as DNA chain terminators. The classical chain-termination method requires 1- single-stranded DNA template 2- DNA primer 3- DNA polymerase 4- normal deoxynucleotidephosphates (dNTPs) 5-modified nucleotides (dideoxyNTPs) that terminate DNA strand elongation. These ddNTPs will also be radioactively or fluorescently labeled for detection in automated sequencing machines.
- 159. In a synthesis reaction, if a dideoxynucleotide is added instead of the normal deoxynucleotide, the synthesis stops at that point because the 3’OH necessary for the addition of the next nucleotide is absent.
- 160. In the dideoxy method of sequencing, the template DNA that is to be sequenced is mixed with a primer complementary to the template DNA and the four normal dNTPs, one of which is radioactively labeled for subsequent visualization purposes. This mixture is then splint into four different tubes that are labeled A, C, G, and T. Each tube is then ―spiked‖ with a different ddNTP (ddATP for tube A, ddCTP for tube C, ddGTT for tube G, or ddTTP for tube T). DNA polymerase is added and using the DNA template and its’ complementary primer, the synthesis of new strands of DNA complementary to the template begins. Occasionally a dideoxynucleotide is added instead of the normal deoxynucleotide and synthesis of that strand is terminated at that point.
- 161. In the tube containing ddATP, some percentage of newly synthesized molecules will get a ddATP in each place that there is a T in the template DNA. The result is a set of new DNA molecules in tube A, each of which ends in an A. A similar type of reaction occurs in the three other tubes to result in molecules that end in C, G, and T in tubes C, G, and T respectively. After the synthesis reactions are complete, the products of the four different tubes are loaded onto four adjacent lane of a polyacrylamide gel and the different fragments are separated by size. The sequencing gel is able to resolve fragments that differ in size from each other by only one base.
- 162. After electrophoresis to separate the fragments by size, the fragments are visualized to exposing the gel to photographic film (Remember that one nucleotide was radioactively labeled). All fragments in lane A will end in an A, fragments in lane C will all end in a C, fragments in lane G will all end in a G, and fragments in lane T will all end in a T. The sequence of the DNA is read from the gel by starting at the bottom and reading upward.
- 164. DNA sequencing: the Sanger method Four separate polymerization reactions are performed
- 165. DNA sequencing: the Sanger (dideoxy) method
- 167. Dye-terminator sequencing Automated DNA sequencing – in automated DNA sequencing a radioactive deoxynucleotide is not used and all four dideoxy reactions are done in a single tube. This is possible because each ddNTPs is labeled with a different flourescent dye. Therefore the dye present in each synthesized fragment corresponds to the dye attached to the dideoxynucleotide that was added to terminate the synthesis of that particular fragment. The contents of the single tube reaction are loaded onto a single lane of a gel and electrophoresis is done.
- 168. A flourimeter and computer are hooked up to the gel and they detect and record the dye attached to the fragments as they come off the gel. The sequence is determined by the order of the dyes coming off the gel.
- 169. Semi-Automated Sequencing •thermal cycler •fluorescent ddNTPs •unique spectra •measure intensity of DNA products on gel
- 172. Automated DNA Sequencing with Fluorescent Dyes Each different ddNTP is coupled to a different colored fluorescent dye ddTTP is red; ddGTP is black etc.
