Somatic variant detection from circulating tumor cells using targeted next-generation sequencing
•
1 like•1,274 views
Somatic variant detection in oncology research may provide insight into drug response and helps identify new therapeutic targets in the future. A key limitation to this process is the availability of tumor tissue that adequately represents the current disease status. Development of a robust method for detecting mutational status from a blood sample would enable analysis of tumor changes over time, which is impractical using tissue samples. A workflow is presented for analyzing somatic variants in circulating tumor cell (CTC) samples using next-generation sequencing (NGS).
Report
Share
Report
Share
Download to read offline
Recommended
The use of cell-free circulating tumor DNA (ctDNA) for non-invasive cancer testing has the potential to revolutionize the field. However, emergence of an increasing number of extraction methods and detection assays is rendering laboratory workflow development much more complex and cumbersome. The use of standardized, well characterized ctDNA control materials in human plasma could facilitate the evaluation of extraction efficiency and assay performance across platforms. In this study, we use a full process ctDNA quality control material in true human plasma to demonstrate the variability of extraction yield between different ctDNA extraction kits. We also examine the correlation between the amplifiable
copy number and DNA concentration post-extraction.Evaluation of ctDNA extraction methods and amplifiable copy number yield usin...

Evaluation of ctDNA extraction methods and amplifiable copy number yield usin...Thermo Fisher Scientific
Recommended
The use of cell-free circulating tumor DNA (ctDNA) for non-invasive cancer testing has the potential to revolutionize the field. However, emergence of an increasing number of extraction methods and detection assays is rendering laboratory workflow development much more complex and cumbersome. The use of standardized, well characterized ctDNA control materials in human plasma could facilitate the evaluation of extraction efficiency and assay performance across platforms. In this study, we use a full process ctDNA quality control material in true human plasma to demonstrate the variability of extraction yield between different ctDNA extraction kits. We also examine the correlation between the amplifiable
copy number and DNA concentration post-extraction.Evaluation of ctDNA extraction methods and amplifiable copy number yield usin...

Evaluation of ctDNA extraction methods and amplifiable copy number yield usin...Thermo Fisher Scientific
Presented here is an analytical validation of OCAv3 at the Life Technologies Clinical Services Laboratory (LTCSL), a CAP-accredited and CLIA-certified clinical laboratory. Analytical validations provide evidence of consistently accurate and relevant sequencing results.Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...

Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...Thermo Fisher Scientific
Accurate identification of uropathogens in a timely manner is important to correctly understand urinary tract infections(UTI’s), which affects nearly 150 million people each year. The
current standard approach for detecting the UTI pathogens is culture based. This method is time consuming, has low throughput, and can lack sensitivity and/or specificity. In addition, not all uropathogens grow equally well under standard culture conditions which can result in a failure to detect the species. To address these gaps, we have developed a unique workflow from sample preparation to target identification using the nanofluidic OpenArray™ platform for spatial multiplexing of target specific assays. In this study, we tested pre-determined blinded research samples and confirmed the subset of results with orthogonal Sanger sequences.Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...

Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...Thermo Fisher Scientific
Liquid biopsy is emerging as a non-invasive companion to traditional solid tumor biopsies. As next generation sequencing (NGS) of circulating cell-free nucleic acids (cfNA = cfDNA and cfRNA) becomes common, it’s important to understand the impact of sample preparation on quality, specificity, and sensitivity of liquid biopsy tests. Plasma samples are often limited, and may have undesirable characteristics such as lipemia or hemolysis that contribute unwanted genomic DNA (gDNA) to the sample. Low cfDNA concentration can also limit the amount available for NGS library prep. In this study, we explore the effects of suboptimal plasma and low library input on liquid biopsy NGS, and discuss various techniques for in-process quality control of cfNA samples isolated from plasmaLiquid biopsy quality control – the importance of plasma quality, sample prep...

Liquid biopsy quality control – the importance of plasma quality, sample prep...Thermo Fisher Scientific
Next generation sequencing (NGS) assay development for solid tumor sequencing requires characterization of variant calling directly from formalin-fixed paraffin embedded (FFPE) tissue samples. However, cell line based FFPE and human FFPE samples only contain 2 to 20 variants, which require laboratories to invest significant resources in sample sourcing and preparation when developing assays to detect 100+ variantsStreamlined next generation sequencing assay development using a highly multi...

Streamlined next generation sequencing assay development using a highly multi...Thermo Fisher Scientific
T-cell receptor beta (TCRβ) immune repertoire analysis by next-generation sequencing is a valuable tool for studies of the tumor microenvironment and potential immune responses to cancer immunotherapy. Here we describe a TCRβ sequencing assay that leverages the low sample input requirements of AmpliSeq library preparation technology to extend the capability of targeted immune repertoire sequencing to include FFPE samples which can often be degraded and in short supplyTargeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...

Targeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...Thermo Fisher Scientific
Targeted next-generation sequencing (NGS) panels can detect hundreds of mutations in key genes using amplification based and hybrid-capture based NGS technologies. Although NGS technology is a powerful tool, optimizing and characterizing test performance on hundreds of variants is extremely challenging, time consuming, and expensive. Samples must be sourced, variants identified and orthogonally confirmed, then quantified and diluted. This effort is then multiplied across dozens of samples, and then samples must be run over many runs and days to assess assay reproducibility, precision, sensitivity, etc. In this study, we developed a novel reference material, experimental design, and analysis pipeline that allows for highly streamlined NGS assay characterization, enabling thorough test characterization across 500+ variants within only 6 runs.Development of Quality Control Materials for Characterization of Comprehensiv...

Development of Quality Control Materials for Characterization of Comprehensiv...Thermo Fisher Scientific
The proliferation of genetic testing technologies and genome-scale studies has increased our understanding of the genetic basis of complex diseases. However, this information alone tells an incomplete story of the underlying biology. Integrative approaches that combine data from multiple sources, such as the genome, transcriptome and/or proteome, can provide a more comprehensive and multi-dimensional model of complex diseases. Similarly, the integration of multiple data types in disease screening can improve our understanding of disease in populations. In a series of groundbreaking multi-omic, population-based studies of prostate cancer, researchers at the Karolinska Institutet in Stockholm, Sweden identified sets of genetic and protein biomarkers that when evaluated together with other clinical research data performed significantly better in predicting cancer risk (1,2) than the most-widely used single protein biomarker, the prostate-specific antigen (PSA).A high-throughput approach for multi-omic testing for prostate cancer research

A high-throughput approach for multi-omic testing for prostate cancer researchThermo Fisher Scientific
Herein, we describe a new research method for library
preparation using the Ion AmpliSeq™ HD Library Kit with
custom assay designs from Ion AmpliSeq HD Panels for
detection of low level variants from liquid biopsy samples. This
method includes incorporation of molecular tags that enable
0.1% Limit of Detection (LOD) in cell free DNA (cfDNA) and
dual barcodes for sample identification. This method is also
applicable to formalin-fixed paraffin embedded (FFPE)
samples. The libraries can be prepared in as little as 3 hours
and are compatible for analysis with the Ion GeneStudio™ S5
systemA rapid library preparation method with custom assay designs for detection of...

A rapid library preparation method with custom assay designs for detection of...Thermo Fisher Scientific
Reprogramming permits the derivation of hiPSCs from diseased patients, and allows us to model diseases in vitro. Furthermore, with the advent of CRISPR mediated genome editing, we can now mimic disease mutations in control hiPSC lines to study the biological effect of just those mutations. hiPSCs can then be differentiated into specified cell types such as neurons which can be used to develop assays for drug safety screening or can be used to model disease phenotypes in a dish to discover new drugs.Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...

Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...Thermo Fisher Scientific
MicroRNAs (miRNA) are a class of small non-coding RNAs (approximately 21 nt long) that bind complementary sequences in target mRNAs to specifically regulate gene expression. Aberrant regulation of miRNAs and their targets has been associated with several diseases including cancer. The relationship between miRNA and mRNA has been found to be important in cancer development and progression. Simultaneous expression studies of miRNA and mRNA and detection of mutations in mRNA transcripts can be valuable in understanding molecular mechanisms that
have an underlying role in various diseases. We demonstrate the technical verification of a novel method to reverse-transcribe and pre-amplify miRNA and mRNA from sample-limiting serum research samples using the TaqMan® Advanced miRNA cDNA Synthesis Kit. Based on results from previous studies, a signature of 49 mRNA and 37 miRNA targets has been identified that may help distinguish between benign and malignant pancreatic tissues. In this study, these targets and an additional set of transcript mutations were analyzed in serum from normal and test samples. TaqMan assays for miRNA and mRNA targets and custom TaqMan Mutation Detection Assays (TMDAs) were placed on TaqMan Array Cards to facilitate investigation of several samples in a single experiment. Results demonstrate that transcript mutations can be detected and miRNA and mRNA targets can be reliably quantified from a single reverse transcription reaction. For research use only. Not for use in diagnostic purposes.TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...

TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...Thermo Fisher Scientific
In this study, we developed a CRISPR/Cas9-based high throughput loss-of-function screen for identifying target genes responsible for the tumor proliferation and growth in TNBC. Our initial focus was to identify essential kinases in MDA-MB-231 cell line using the Invitrogen™ LentiArray™ Human Kinase CRISPR Library, which targets 840 kinases with up to 4 different gRNAs per protein kinase for complete gene knockout. This functional screen identified over 90 protein kinases that are essential for cell viability and cell proliferation. Ten of these hits (CDK1, CDK2, CDK8, CDK10, CDK11A, CDK19, CDK19, CDC7, EPHA2 and WEE1) are well-known targets validated in the literature. Currently, we are in the process validating the novel hits through target gene sequencing, western blotting and target specific small molecule kinase inhibitors.Identifying novel and druggable targets in a triple negative breast cancer ce...

Identifying novel and druggable targets in a triple negative breast cancer ce...Thermo Fisher Scientific
T cell convergence refers to the phenomenon whereby antigen-driven selection enriches for T cell receptors (TCRs) having a shared antigen specificity but different amino acid or
nucleotide sequence. T cell recruitment and expansion within the tumor microenvironment (TME) may be directed by responses to tumor neoantigen, suggesting that elevated T
cell convergence could be a general feature of the tumor infiltrating T cell repertoire. Here we use the Ion AmpliSeq™ Immune Repertoire Assay Plus – TCRβ to evaluate evidence
for T cell convergence within melanoma tumor biopsy research samples from a set of 63 subjects plus peripheral blood leukocytes (PBL) from four healthy subjects. We find that the melanoma TME is highly enriched for convergent TCRs compared to healthy donor peripheral blood. We discuss the potential use of TCR convergence as a liquid biopsy compatible predictive biomarker for immunotherapy response.Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...

Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...Thermo Fisher Scientific
To support clinical and translational research into precision oncology strategies for myeloid cancers, a next-generation sequencing (NGS) assay was developed to detect common and relevant somatic alterations. To define gene targets that were recurrently altered in myeloid cancers and relevant for clinical and translational research, an extensive survey of investigators at hematology oncology research labs was performed.Analytical performance of a novel next generation sequencing assay for Myeloi...

Analytical performance of a novel next generation sequencing assay for Myeloi...Thermo Fisher Scientific
Tumor mutation load predicts durable benefit from immune checkpoint inhibitors in several cancer types. Existing methods to estimate tumor mutation load have large input DNA and extensive infrastructure requirements and are associated with delays due to shipping biopsy samples to central laboratories. We demonstrate the ability of a targeted panel with fast
turn-around time and low input requirement for estimating mutation load from tumor samples to advance research in immuno-oncology.Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...

Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...Thermo Fisher Scientific
The study of recurrent somatic alterations associated with pediatric, childhood and young adult cancers has lagged behind those that associated with adult cancers. Whole exome and transcriptome approaches are still being used to support discovery efforts, consequently, due to several initiatives aimed at profiling genomic alterations associated with childhood cancers, a set of recurrent somatic alterations has been defined.Development of a next-generation (NGS) assay for pediatric, childhood, and yo...

Development of a next-generation (NGS) assay for pediatric, childhood, and yo...Thermo Fisher Scientific
Oxygen levels in typical cell culture conditions do not accurately reflect the oxygen levels cells are exposed to within the body. Furthermore, oxygen levels can vary within the tumor microenvironment. These variances can affect how cells respond to a variety of drugs and small molecules. To further understand how oxygen levels affect drug sensitivity, the response of hormone-dependent MCF7 cells were compared to hormone-independent MDA-MB231 cells, cultured under low and high oxygen.High content screening in MCF7 and MDA-MB231 cells show differential response...

High content screening in MCF7 and MDA-MB231 cells show differential response...Thermo Fisher Scientific
Pharmacogenomics (PGx) is the study of genetic variations in terms of their response to drugs. Variations in gene sequence or copy numbers may result in complete loss of function, partial decrease or increase in enzyme activity, or an altered affinity for substrates, which may in turn significantly impact drug efficacy. PGx studies are becoming increasingly important for precision medicine. We have developed a next generation sequencing (NGS) PGx research solution with increased flexibility on the assay targets and combined detection of SNP/INDEL genotyping and CNV using Ion AmpliSeq™ technology for low to medium throughput laboratories. With this highly multiplexed PGx research panel we can profile a set of 136 genetic markers in 40 known PGx related genes (Table 1) and determine CYP2D6 copy number variation (CNV, Figure 1) in a single reaction using Ion Torrent™ semiconductor sequencing.A next generation sequencing based sample-to-result pharmacogenomics research...

A next generation sequencing based sample-to-result pharmacogenomics research...Thermo Fisher Scientific
More Related Content
More from Thermo Fisher Scientific
Presented here is an analytical validation of OCAv3 at the Life Technologies Clinical Services Laboratory (LTCSL), a CAP-accredited and CLIA-certified clinical laboratory. Analytical validations provide evidence of consistently accurate and relevant sequencing results.Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...

Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...Thermo Fisher Scientific
Accurate identification of uropathogens in a timely manner is important to correctly understand urinary tract infections(UTI’s), which affects nearly 150 million people each year. The
current standard approach for detecting the UTI pathogens is culture based. This method is time consuming, has low throughput, and can lack sensitivity and/or specificity. In addition, not all uropathogens grow equally well under standard culture conditions which can result in a failure to detect the species. To address these gaps, we have developed a unique workflow from sample preparation to target identification using the nanofluidic OpenArray™ platform for spatial multiplexing of target specific assays. In this study, we tested pre-determined blinded research samples and confirmed the subset of results with orthogonal Sanger sequences.Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...

Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...Thermo Fisher Scientific
Liquid biopsy is emerging as a non-invasive companion to traditional solid tumor biopsies. As next generation sequencing (NGS) of circulating cell-free nucleic acids (cfNA = cfDNA and cfRNA) becomes common, it’s important to understand the impact of sample preparation on quality, specificity, and sensitivity of liquid biopsy tests. Plasma samples are often limited, and may have undesirable characteristics such as lipemia or hemolysis that contribute unwanted genomic DNA (gDNA) to the sample. Low cfDNA concentration can also limit the amount available for NGS library prep. In this study, we explore the effects of suboptimal plasma and low library input on liquid biopsy NGS, and discuss various techniques for in-process quality control of cfNA samples isolated from plasmaLiquid biopsy quality control – the importance of plasma quality, sample prep...

Liquid biopsy quality control – the importance of plasma quality, sample prep...Thermo Fisher Scientific
Next generation sequencing (NGS) assay development for solid tumor sequencing requires characterization of variant calling directly from formalin-fixed paraffin embedded (FFPE) tissue samples. However, cell line based FFPE and human FFPE samples only contain 2 to 20 variants, which require laboratories to invest significant resources in sample sourcing and preparation when developing assays to detect 100+ variantsStreamlined next generation sequencing assay development using a highly multi...

Streamlined next generation sequencing assay development using a highly multi...Thermo Fisher Scientific
T-cell receptor beta (TCRβ) immune repertoire analysis by next-generation sequencing is a valuable tool for studies of the tumor microenvironment and potential immune responses to cancer immunotherapy. Here we describe a TCRβ sequencing assay that leverages the low sample input requirements of AmpliSeq library preparation technology to extend the capability of targeted immune repertoire sequencing to include FFPE samples which can often be degraded and in short supplyTargeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...

Targeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...Thermo Fisher Scientific
Targeted next-generation sequencing (NGS) panels can detect hundreds of mutations in key genes using amplification based and hybrid-capture based NGS technologies. Although NGS technology is a powerful tool, optimizing and characterizing test performance on hundreds of variants is extremely challenging, time consuming, and expensive. Samples must be sourced, variants identified and orthogonally confirmed, then quantified and diluted. This effort is then multiplied across dozens of samples, and then samples must be run over many runs and days to assess assay reproducibility, precision, sensitivity, etc. In this study, we developed a novel reference material, experimental design, and analysis pipeline that allows for highly streamlined NGS assay characterization, enabling thorough test characterization across 500+ variants within only 6 runs.Development of Quality Control Materials for Characterization of Comprehensiv...

Development of Quality Control Materials for Characterization of Comprehensiv...Thermo Fisher Scientific
The proliferation of genetic testing technologies and genome-scale studies has increased our understanding of the genetic basis of complex diseases. However, this information alone tells an incomplete story of the underlying biology. Integrative approaches that combine data from multiple sources, such as the genome, transcriptome and/or proteome, can provide a more comprehensive and multi-dimensional model of complex diseases. Similarly, the integration of multiple data types in disease screening can improve our understanding of disease in populations. In a series of groundbreaking multi-omic, population-based studies of prostate cancer, researchers at the Karolinska Institutet in Stockholm, Sweden identified sets of genetic and protein biomarkers that when evaluated together with other clinical research data performed significantly better in predicting cancer risk (1,2) than the most-widely used single protein biomarker, the prostate-specific antigen (PSA).A high-throughput approach for multi-omic testing for prostate cancer research

A high-throughput approach for multi-omic testing for prostate cancer researchThermo Fisher Scientific
Herein, we describe a new research method for library
preparation using the Ion AmpliSeq™ HD Library Kit with
custom assay designs from Ion AmpliSeq HD Panels for
detection of low level variants from liquid biopsy samples. This
method includes incorporation of molecular tags that enable
0.1% Limit of Detection (LOD) in cell free DNA (cfDNA) and
dual barcodes for sample identification. This method is also
applicable to formalin-fixed paraffin embedded (FFPE)
samples. The libraries can be prepared in as little as 3 hours
and are compatible for analysis with the Ion GeneStudio™ S5
systemA rapid library preparation method with custom assay designs for detection of...

A rapid library preparation method with custom assay designs for detection of...Thermo Fisher Scientific
Reprogramming permits the derivation of hiPSCs from diseased patients, and allows us to model diseases in vitro. Furthermore, with the advent of CRISPR mediated genome editing, we can now mimic disease mutations in control hiPSC lines to study the biological effect of just those mutations. hiPSCs can then be differentiated into specified cell types such as neurons which can be used to develop assays for drug safety screening or can be used to model disease phenotypes in a dish to discover new drugs.Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...

Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...Thermo Fisher Scientific
MicroRNAs (miRNA) are a class of small non-coding RNAs (approximately 21 nt long) that bind complementary sequences in target mRNAs to specifically regulate gene expression. Aberrant regulation of miRNAs and their targets has been associated with several diseases including cancer. The relationship between miRNA and mRNA has been found to be important in cancer development and progression. Simultaneous expression studies of miRNA and mRNA and detection of mutations in mRNA transcripts can be valuable in understanding molecular mechanisms that
have an underlying role in various diseases. We demonstrate the technical verification of a novel method to reverse-transcribe and pre-amplify miRNA and mRNA from sample-limiting serum research samples using the TaqMan® Advanced miRNA cDNA Synthesis Kit. Based on results from previous studies, a signature of 49 mRNA and 37 miRNA targets has been identified that may help distinguish between benign and malignant pancreatic tissues. In this study, these targets and an additional set of transcript mutations were analyzed in serum from normal and test samples. TaqMan assays for miRNA and mRNA targets and custom TaqMan Mutation Detection Assays (TMDAs) were placed on TaqMan Array Cards to facilitate investigation of several samples in a single experiment. Results demonstrate that transcript mutations can be detected and miRNA and mRNA targets can be reliably quantified from a single reverse transcription reaction. For research use only. Not for use in diagnostic purposes.TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...

TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...Thermo Fisher Scientific
In this study, we developed a CRISPR/Cas9-based high throughput loss-of-function screen for identifying target genes responsible for the tumor proliferation and growth in TNBC. Our initial focus was to identify essential kinases in MDA-MB-231 cell line using the Invitrogen™ LentiArray™ Human Kinase CRISPR Library, which targets 840 kinases with up to 4 different gRNAs per protein kinase for complete gene knockout. This functional screen identified over 90 protein kinases that are essential for cell viability and cell proliferation. Ten of these hits (CDK1, CDK2, CDK8, CDK10, CDK11A, CDK19, CDK19, CDC7, EPHA2 and WEE1) are well-known targets validated in the literature. Currently, we are in the process validating the novel hits through target gene sequencing, western blotting and target specific small molecule kinase inhibitors.Identifying novel and druggable targets in a triple negative breast cancer ce...

Identifying novel and druggable targets in a triple negative breast cancer ce...Thermo Fisher Scientific
T cell convergence refers to the phenomenon whereby antigen-driven selection enriches for T cell receptors (TCRs) having a shared antigen specificity but different amino acid or
nucleotide sequence. T cell recruitment and expansion within the tumor microenvironment (TME) may be directed by responses to tumor neoantigen, suggesting that elevated T
cell convergence could be a general feature of the tumor infiltrating T cell repertoire. Here we use the Ion AmpliSeq™ Immune Repertoire Assay Plus – TCRβ to evaluate evidence
for T cell convergence within melanoma tumor biopsy research samples from a set of 63 subjects plus peripheral blood leukocytes (PBL) from four healthy subjects. We find that the melanoma TME is highly enriched for convergent TCRs compared to healthy donor peripheral blood. We discuss the potential use of TCR convergence as a liquid biopsy compatible predictive biomarker for immunotherapy response.Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...

Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...Thermo Fisher Scientific
To support clinical and translational research into precision oncology strategies for myeloid cancers, a next-generation sequencing (NGS) assay was developed to detect common and relevant somatic alterations. To define gene targets that were recurrently altered in myeloid cancers and relevant for clinical and translational research, an extensive survey of investigators at hematology oncology research labs was performed.Analytical performance of a novel next generation sequencing assay for Myeloi...

Analytical performance of a novel next generation sequencing assay for Myeloi...Thermo Fisher Scientific
Tumor mutation load predicts durable benefit from immune checkpoint inhibitors in several cancer types. Existing methods to estimate tumor mutation load have large input DNA and extensive infrastructure requirements and are associated with delays due to shipping biopsy samples to central laboratories. We demonstrate the ability of a targeted panel with fast
turn-around time and low input requirement for estimating mutation load from tumor samples to advance research in immuno-oncology.Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...

Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...Thermo Fisher Scientific
The study of recurrent somatic alterations associated with pediatric, childhood and young adult cancers has lagged behind those that associated with adult cancers. Whole exome and transcriptome approaches are still being used to support discovery efforts, consequently, due to several initiatives aimed at profiling genomic alterations associated with childhood cancers, a set of recurrent somatic alterations has been defined.Development of a next-generation (NGS) assay for pediatric, childhood, and yo...

Development of a next-generation (NGS) assay for pediatric, childhood, and yo...Thermo Fisher Scientific
Oxygen levels in typical cell culture conditions do not accurately reflect the oxygen levels cells are exposed to within the body. Furthermore, oxygen levels can vary within the tumor microenvironment. These variances can affect how cells respond to a variety of drugs and small molecules. To further understand how oxygen levels affect drug sensitivity, the response of hormone-dependent MCF7 cells were compared to hormone-independent MDA-MB231 cells, cultured under low and high oxygen.High content screening in MCF7 and MDA-MB231 cells show differential response...

High content screening in MCF7 and MDA-MB231 cells show differential response...Thermo Fisher Scientific
Pharmacogenomics (PGx) is the study of genetic variations in terms of their response to drugs. Variations in gene sequence or copy numbers may result in complete loss of function, partial decrease or increase in enzyme activity, or an altered affinity for substrates, which may in turn significantly impact drug efficacy. PGx studies are becoming increasingly important for precision medicine. We have developed a next generation sequencing (NGS) PGx research solution with increased flexibility on the assay targets and combined detection of SNP/INDEL genotyping and CNV using Ion AmpliSeq™ technology for low to medium throughput laboratories. With this highly multiplexed PGx research panel we can profile a set of 136 genetic markers in 40 known PGx related genes (Table 1) and determine CYP2D6 copy number variation (CNV, Figure 1) in a single reaction using Ion Torrent™ semiconductor sequencing.A next generation sequencing based sample-to-result pharmacogenomics research...

A next generation sequencing based sample-to-result pharmacogenomics research...Thermo Fisher Scientific
More from Thermo Fisher Scientific (20)
Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...

Analytical Validation of the Oncomine™ Comprehensive Assay v3 with FFPE and C...
Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...

Novel Spatial Multiplex Screening of Uropathogens Associated with Urinary Tra...
Liquid biopsy quality control – the importance of plasma quality, sample prep...

Liquid biopsy quality control – the importance of plasma quality, sample prep...
Streamlined next generation sequencing assay development using a highly multi...

Streamlined next generation sequencing assay development using a highly multi...
Targeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...

Targeted T-cell receptor beta immune repertoire sequencing in several FFPE ti...
Development of Quality Control Materials for Characterization of Comprehensiv...

Development of Quality Control Materials for Characterization of Comprehensiv...
A High Throughput System for Profiling Respiratory Tract Microbiota

A High Throughput System for Profiling Respiratory Tract Microbiota
A high-throughput approach for multi-omic testing for prostate cancer research

A high-throughput approach for multi-omic testing for prostate cancer research
Why is selecting the right thermal cycler important?

Why is selecting the right thermal cycler important?
A rapid library preparation method with custom assay designs for detection of...

A rapid library preparation method with custom assay designs for detection of...
Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...

Generation of Clonal CRISPR/Cas9-edited Human iPSC Derived Cellular Models an...
TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...

TaqMan®Advanced miRNA cDNA synthesis kit to simultaneously study expression o...
Identifying novel and druggable targets in a triple negative breast cancer ce...

Identifying novel and druggable targets in a triple negative breast cancer ce...
Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...

Evidence for antigen-driven TCRβ chain convergence in the melanoma-infiltrati...
Analytical performance of a novel next generation sequencing assay for Myeloi...

Analytical performance of a novel next generation sequencing assay for Myeloi...
Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...

Estimating Mutation Load from Tumor Research Samples using a Targeted Next-Ge...
Development of a next-generation (NGS) assay for pediatric, childhood, and yo...

Development of a next-generation (NGS) assay for pediatric, childhood, and yo...
High content screening in MCF7 and MDA-MB231 cells show differential response...

High content screening in MCF7 and MDA-MB231 cells show differential response...
A next generation sequencing based sample-to-result pharmacogenomics research...

A next generation sequencing based sample-to-result pharmacogenomics research...
A Comprehensive Childhood Cancer Research Gene Panel

A Comprehensive Childhood Cancer Research Gene Panel
Recently uploaded
Recently uploaded (20)
Chemical Tests; flame test, positive and negative ions test Edexcel Internati...

Chemical Tests; flame test, positive and negative ions test Edexcel Internati...
Pulmonary drug delivery system M.pharm -2nd sem P'ceutics

Pulmonary drug delivery system M.pharm -2nd sem P'ceutics
❤Jammu Kashmir Call Girls 8617697112 Personal Whatsapp Number 💦✅.

❤Jammu Kashmir Call Girls 8617697112 Personal Whatsapp Number 💦✅.
Discovery of an Accretion Streamer and a Slow Wide-angle Outflow around FUOri...

Discovery of an Accretion Streamer and a Slow Wide-angle Outflow around FUOri...
Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b

Asymmetry in the atmosphere of the ultra-hot Jupiter WASP-76 b
Forensic Biology & Its biological significance.pdf

Forensic Biology & Its biological significance.pdf
Formation of low mass protostars and their circumstellar disks

Formation of low mass protostars and their circumstellar disks
High Class Escorts in Hyderabad ₹7.5k Pick Up & Drop With Cash Payment 969456...

High Class Escorts in Hyderabad ₹7.5k Pick Up & Drop With Cash Payment 969456...
Labelling Requirements and Label Claims for Dietary Supplements and Recommend...

Labelling Requirements and Label Claims for Dietary Supplements and Recommend...
SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE

SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE
Connaught Place, Delhi Call girls :8448380779 Model Escorts | 100% verified

Connaught Place, Delhi Call girls :8448380779 Model Escorts | 100% verified
Somatic variant detection from circulating tumor cells using targeted next-generation sequencing
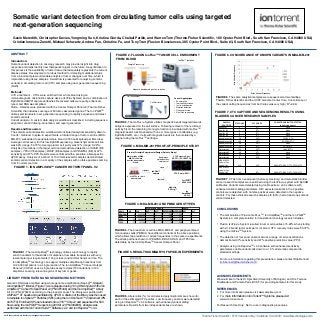
- 1. Gavin Meredith, Christopher Davies, Yongming Sun, Kristina Giorda, Chaitali Parikh, and Warren Tom (Thermo Fisher Scientific, 180 Oyster Point Blvd., South San Francisco, CA 94080 USA) Cristian Ionescu-Zanetti, Michael Schwartz, Andrea Fan, Christine Fu, and Tony Tran (Fluxion Biosciences, 385 Oyster Point Blvd., Suite #3, South San Francisco, CA 94080 USA) FIGURE 2. FLUXION IsoFlux™ TUMOR CELL ENRICHMENT FROM BLOOD ABSTRACT Introduction: Somatic variant detection in oncology research may provide insight into drug response and helps identify new therapeutic targets in the future. A key limitation to this process is the availability of tumor tissue that adequately represents the current disease status. Development of a robust method for detecting mutational status from a blood sample would enable analysis of tumor changes over time, which is impractical using tissue samples. A workflow is presented for analyzing somatic variants in circulating tumor cell (CTC) samples using next-generation sequencing (NGS). Methods: CTC enrichment - CTCs were enriched from whole blood using an immunomagnetic microfluidic capture device (IsoFlux System) and a combination of EpCAM and EGFR capture antibodies. Recovered cells were purity-enhanced, lysed, and DNA was amplified. NGS – Libraries were produced with the Cancer Hotspot Panel v2 (Thermo Fisher Scientific) that includes coverage of 50 tumor-associated genes. The Ion PGMTM utilizes semiconductor next-generation sequencing to rapidly sequence and detect somatic variants. Variant analysis - A custom data analysis workflow is described, including sequence alignment, variant filtering, annotation, and report generation. Results and Discussion: The somatic variant detection workflow demonstrated analytical sensitivity down to 3 cells / mLblood. In spike-in experiments, a titrated range of tumor cell line (MDA- MB-231) was added to peripheral whole blood (0-180 cells spiked per 8mL tube). Average cell recovery for this low-EpCAM expressing, mesenchymal-like cell line was 66% (range: 54-73%). Average tumor cell purity was 12% (range: 5-22%). Analysis of mutations in the target cell-line demonstrated detection of KRAS (8/8 samples), TP53 (7/8 samples), BRAF (8/8 samples) and PDGFRA (8/8) at 97% sensitivity and 97% PPV. Results were confirmed with a sensitive, allele-specific qPCR assay. Analysis of a cohort of 10 clinical research samples demonstrated somatic variant detection in a majority of the samples with no false positives coming from 5 normal controls. CONCLUSIONS • The combination of Fluxion IsoFLuxTM, Ion AmpliSeqTM and the Ion PGMTM System is a complete solution for translational oncology research studies. • Fluxion IsoFluxTM System resulted in tumor cell purities of 5-22% when starting with 2-12 breast tumor cells per ml of blood. CTC recovery rates were 53-72% using the IsoFluxTM System. • The detection of four know variants across a range of cell concentrations demonstrated 97% sensitivity and 97% positive prediction value (PPV). • Analysis using Ion ReporterTM v4.4 software with enhanced sensitivity parameters enable variant calls down to a 1.0% allele ratio compared to standard settings. • For more information regarding this presentation, please contact Rob Bennett (rob.bennett@thermofisher.com). ACKNOWLEDGEMENTS We would like to thank Dr. Ajjai Alva (University of Michigan), and Drs. Terence Friedlander and Pamela Paris (UCSF) for providing samples for this study. REFERENCES 1. For more information please visit www.ampliseq.com 2. For more information on Ion Torrent™ Systems, please visit www.iontorrent.com For Research Use Only. Not for use in diagnostic procedures. Somatic variant detection from circulating tumor cells using targeted next-generation sequencing Thermo Fisher Scientific • 5791 Van Allen Way • Carlsbad, CA 92008 • www.lifetechnologies.comFor Research Use Only. Not for use in diagnostic procedures. FIGURE 7. CTC CAPTURE AND SEQUENCING RESULTS USING BLADDER CANCER RESEARCH SAMPLES FIGURE 7. CTCs from neoadjuvant (before cystectomy) and metastatic bladder cancer research samples were enriched using the IsoFluxTM system with EpCAM antibodies. Variants were detected using Ion ReporterTM v4.4 software with enhanced variant calling parameters. All 3 expected variants in the 2 positive controls were detected, with no false positives were detected in the negative controls. For these clinical research samples, 4/8 (50%) had at least one somatic variant detected. LIBRARY PREPARATION AND SEQUENCING METHODS Genomic DNA was amplified using whole genome amplification (Repli-g™, Qiagen). Ion AmpliSeq™ libraries (Figure 1) were prepared using 10 ng DNA input with the Ion AmpliSeq™ Cancer Hotspot Panel v2 (PN 4475346) and the Ion AmpliSeq™ Library Kit 2.0 (PN 4475345) according to the product manual. The library was purified with AMPure™ XT beads and quantitated by qPCR. A dilution of the library was then used to template Ion Sphere™ Particles (ISPs) using the Ion OneTouch™ 2 instrument (PN 4474779). Enriched ISPs were loaded onto a 318™ Chip v2 and sequenced for 500 flows using the Ion PGM™ Sequencing 200 Kit v2 (PN 4482006). Analysis was performed with the Ion Torrent Suite™ Software v.4.2 and Ion Reporter™ v4.4. SAMPLE IDENTIFIER CTC COUNT CTC PURITY SOMATIC VARIANTS DETECTED (allelic frequency) Healthy #1 0 0% None Healthy #2 0 0% None Healthy #3 0 0% None Healthy #4 0 0% None Neoadjuvant #1 40 10% FGFR2 (1%) Neoadjuvant #2 100 19% PDGFRA (1%) Neoadjuvant #3 20 12% None Neoadjuvant #4 21 8% EGFR (6%) Metastatic #1 154 25% JAK2 (12%) Metastatic #2 34 4% None Metastatic #3 11 15% None Metastatic #4 8 N/D None MDA-MB-231 Spike In (Pos Control) #1 N/D N/D KRAS (6%), BRAF (7%), TP53 (12%) MDA-MB-231 Spike In (Pos Control) #2 60 8% KRAS (4%), BRAF (6%), TP53 (11%) Proof of Principle Experimental Design (Spike-In Study) Mixture study of MDA- MB-231 cells into healthy donor whole blood triplicate IsoFlux™ capture 0 - 314 CTCs in 14 mL whole blood Enumerate spiked cells vs. total nuclei CTC Recovery Rates: 53-72% (32-169 CTCs) CTC Purity: 5-22% single WGA with REPLI-g™ duplicate Cast-PCR™ KRAS G13D Ion AmpliSeq™ CHPv2 Sample Processing Cell Enrichment Cell Retrieval IsoFlux™ Isolation Magnetic bead based positive selection Current Applications • Enumeration • Immunofluorescence • In situ hybridization • Mutation detection • Gene expression • Next Gen sequencing FIGURE 1. ION AMPLISEQ™ CANCER HOTSPOT PANEL V2 FIGURE 1. The Ion AmpliSeqTM technology utilizes a technology to rapidly enrich hundreds to thousands of mutations at low allele frequencies with very low sample input requirements (10ng or less) and minlmal hands on time. The Ion AmpliSeqTM technology can support multiplex amplification reactions from 12-24,000 amplicons in a single reaction. The Ion AmpliSeqTM Cancer Hotspot Panel v2 (CHPv2) uses a single-tube assay to detect 739 mutations in 190 amplicons covering relevant regions of key tumor genes. FIGURE 2. The IsoFluxTM System utilizes magnetic beads targeted towards antigens expressed on the cell surface. Following removal of the red blood cells by Ficoll, the remaining monocyte fraction is incubated with IsoFluxTM magnetic beads functionalized with one or more types of antibodies (e.g. EpCAM, EGFR, etc.). Cells with magnetic beads are then isolated by a magnet using the IsoFluxTM cartridge. FIGURE 3. MDA-MB-231 PROOF-OF-PRINCIPLE STUDY FIGURE 4. MDA-MB-231 AND PBMC GENOTYPES FIGURE 4. The breast tumor cell line MDA-MB-321 and peripheral blood mononuclear cells (PBMCs) have different variants at the above positions, which allows the prediction of allelic frequencies in the spike-in experiments. The allelic positions shown for PDGFRA, BRAF, KRAS, and TP53 are detectable by the Ion AmpliSeqTM Cancer Hotspot Panel. FIGURE 5. Allele ratios (%) for all detected polymorphisms are shown for each of the estimated CTC purities. Low frequency variants were detected using Ion ReporterTM v4.4 software with enhanced variant calling parameters. Results from two independent sites are shown. FIGURE 5. RESULTS SUMMARY OF SPIKE-IN EXPERIMENTS FIGURE 6. CONCORDANCE OF KNOWN VARIANTS IN MDA-MB-231 Site 1 Variant Frequency Site2VariantFrequency FIGURE 6. The same analytical samples were sequenced at two facilities, Thermo Fisher Scientific and the UCSF Genome Center Core. Concordance of the variant calling frequencies from both sites was very high ( R2=0.94). MDA-MB231 Spike-In PBMA GENE CHR POS REF ALT REF COUNT ALT COUNT REF ALT PDGFRA 4 55152040 C T 2391 1331 C - BRAF 7 140481417 C A 1161 1755 C - KRAS 12 25398281 C T 986 1924 C - TP53 17 7577099 C T 14 2462 C -
