FINAL HHMI POSTER Byanjana Thapa
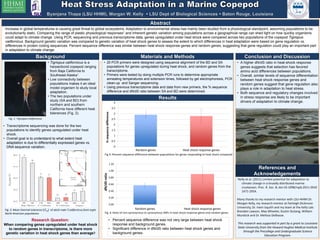
- 1. Abstract Increase in global temperatures is causing great threat to global ecosystems. Adaptation to environmental stress has mainly been studied from a physiological standpoint, assuming populations to be evolutionarily static. Comparing the range of plastic physiological responses1 and inherent genetic variation among populations across a geographical range can shed light on how quickly organisms could adapt to climate change. Using PCR, sequencing and previous transcriptome data, genes upregulated under heat shock were compared across two populations of the copepod Tigriopus californicus. Background genetic variation was compared to genetic variation of heat shock genes to assess the extent to which differences in heat adaptation were based on gene regulation vs. differences in protein coding sequences. Percent sequence difference was similar between heat shock response genes and random genes, suggesting that gene regulation could play an important part in adaptation to climate change. Results Conclusion and Discussion • A higher dN/dS ratio in heat-shock response genes suggests that selection has favored amino acid differences between populations. • Overall, similar levels of sequence differentiation between heat shock response genes and random genes suggest that gene regulation also plays a role in adaptation to heat stress. • Both sequence and regulatory changes involved in stress response are likely to be important drivers of adaptation to climate change. Background • Tigriopus californicus is a harpacticoid copepod ranging from Baja California to Southeast Alaska1. • Low connectivity between populations makes it an ideal model organism to study local adaptation. • The two populations under study (SA and BD) from northern and southern California have different heat tolerances (Fig. 2). • Transcriptome sequencing was done for the two populations to identify genes upregulated under heat shock. • Overall goal is to understand to what extent heat adaptation is due to differentially expressed genes vs. DNA sequence variation. Research Question: When comparing genes upregulated under heat shock to random genes in transcriptome, is there more genetic variation in heat shock genes than average? Materials and Methods • 20 PCR primers were designed using sequence alignment of the BD and SA populations for genes upregulated during heat shock, and random genes from the transcriptome. • Primers were tested by doing multiple PCR runs to determine appropriate annealing temperatures and extension times, followed by gel electrophoresis, PCR clean-up and Sanger sequencing. • Using previous transcriptome data and data from new primers, the % sequence difference and dN/dS ratio between SA and BD were determined. References and Acknowledgements 1Kelly et al. (2011) Limited potential for adaptation to climate change in a broadly distributed marine crustacean. Proc. R. Soc. B; doi:10.1098/rspb.2011.0542 1471-2954. Many thanks to my research mentor with LSU-HHMI Dr. Morgan Kelly, my research mentor at Fairleigh Dickinson University, Dr. Irwin Isquith and my team at the Kelly Lab: Branden Lawson, Max Wheeler, Dustin DuSang, William Murdock and Dr. Melissa DeBiasse. This research was supported in part by a grant to Louisiana State University from the Howard Hughes Medical Institute through the Precollege and Undergraduate Science Education Program. Fig. 1: Tigriopus californicus 0 0.5 1 1.5 2 2.5 3 3.5 4 Random genes Heat shock response genes %sequencedifference Fig 3: Percent sequence difference between populations for genes responding to heat shock compared 0.00 0.20 0.40 0.60 0.80 1.00 1.20 Random genes Heat shock response genes dN/dSratio Fig. 4: Ratio of non-synonymous to synonymous SNPs in heat shock response genes and random genes • Percent sequence difference was not very large between heat shock response and background genes. • Significant difference in dN/dS ratio between heat shock genes and background genes. Fig. 2: Mean thermal tolerance (LT50) of adult male T.californicus from eight North American populations
Hinweis der Redaktion
- Selection has favored some aa differences between populations