RDFizing PubMed Central with Biotea
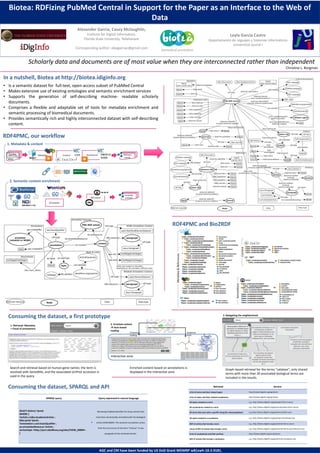
- 1. Biotea: RDFizing PubMed Central in Support for the Paper as an Interface to the Web of Data Alexander Garcia, Casey Mclaughlin, Institute for Digital Information, Florida State University. Tallahassee Leyla Garcia Castro Departamento de Leguajes y Sistemas Informáticos Universitat Jaumé I Corresponding author: alexgarciac@gmail.com Scholarly data and documents are of most value when they are interconnected rather than independent Christine L. Borgman In a nutshell, Biotea at http://biotea.idiginfo.org • Is a semantic dataset for full-text, open-access subset of PubMed Central • Makes extensive use of existing ontologies and semantic enrichment services • Supports the generation of self-describing machine- readable scholarly documents. • Comprises a flexible and adaptable set of tools for metadata enrichment and semantic processing of biomedical documents. • Provides semantically rich and highly interconnected dataset with self-describing content. RDF4PMC, our workflow 1. Metadata & content Metadata Content RDFized article Provenance NXML 2. Semantic content enrichment Enriched content RDFization Annotation RDF4PMC and Bio2RDF Consuming the dataset, a first prototype 3. Navigating the neighborhood 2. Enriched content facts-based reading 1. Retrieval: Metadata + Cloud of annotations Contextual reading Graphical tools Interactive zone Search and retrieval based on human gene names: the term is resolved with GeneWiki, and the associated UniProt accession is used in the query Enriched content based on annotations is displayed in the interactive zone Graph-based retrieval for the terms “catalase”; only shared terms with more than 30 associated biological terms are included in the results. Consuming the dataset, SPARQL and API Retrieval Service A list of terms and their related topics SELECT distinct ?pmid WHERE { ?article a bibo:AcademicArticle ; bibo:pmid ?pmid . ?annotation a aot:ExactQualifier ; ao:annotatesResource ?article ; ao:hasTopic <http://purl.obolibrary.org/obo/CHEBI_60004> . } have been semantically annotated with the biological entity CHEBI:60004. The semantic annotation comes from the occurrence of the term “mixture” in any paragraph of the retrieved articles. e.g., http://biotea.idiginfo.org/api/topics?term=cancer e.g., http://biotea.idiginfo.org/api/vocabularies?term=cancer All terms that start with a specific string (for autocompletion) e.g.,http://biotea.idiginfo.org/api/terms?prefix=canc All topics related to a vocabulary e.g., http://biotea.idiginfo.org/api/topics?vocabulary=po RDF of articles that include a term e.g., http://biotea.idiginfo.org/api/articles?term=cancer Count of RDF of articles that include a term e.g., http://biotea.idiginfo.org/api/articles?term=cancer&count=true A list of vocabularies and their prefixes http://biotea.idiginfo.org/vocabularies RDF of articles that include a vocabulary Retrieving PubMed identifier for those articles that http://biotea.idiginfo.org/api/topics All vocabularies related to a term Query expressed in natural language A list of topics and their related vocabularies All topics related to a term SPARQL query http://biotea.idiginfo.org/api/terms e.g., http://biotea.idiginfo.org/api/articles?vocabulary=po AGC and CM have been funded by US DoD Grant MOMRP w81xwh-10-2-0181.
